Izsvák Lab
Mobile DNA
Transposon-host interaction
Project #3
Elaborate the roles of various HERVH transcripts during human preimplantation embryogenesis
Aleksandra Kondrashkina
HERVH is a primate-specific endogenous retrovirus, which was domesticated during the last 10 million years. ~300 HERVH-derived transcripts have been implicated to be incorporated into the regulatory network of primate/human pluripotency. The goal of the project is to elaborate the roles of various HERVH transcripts during human preimplantation embryogenesis.
Replication stress triggers Sleeping Beauty activity at replication forks
Suneel Narayanavari, Bertrand Tangu Teneng, Andrea Smith*
The vertebrate transposon, Sleeping Beauty (SB) provides an excellent model to investigate host-transposon interactions. During the cut and paste process of transposition, the SB transposon is excised from its original location by the transposes, and is integrated into a new location. SB excision produces a DNA double stranded break (DSB) at the donor site, whereas integration generates single-stranded gaps flanking the element at the insertion site. In mammalian somatic cells, excision sites are primarily repaired by non-homologous end joining (NHEJ), however, in contrast to V(D)J recombination, the utilization of NHEJ is not absolute. Our recent data show that SB-generated excision sites can be accessed by alternative DSB repair (DSBR) pathways. Our SILAC strategy coupled with mass spectrometry revealed that SB is capable of recruiting key DSBR signalling and repair factors of alternative pathways throughout the cell cycle. Pathway choice is determined by the availability of repair factors and their affinity to the transposase. We identified a PIP-like PCNA-binding domain in the SB transposase. Curiously, SB avoids elongating replication forks in normally cycling cells, but binds the forks in the stalled state with an accompanying increase in transposition efficiency. Stalled replication forks comprise extensive single-stranded DNA and elicit a strong checkpoint response. Antagonism of ATR, a replication specific DNA-damage signalling protein impairs SB transposition, suggesting that SB senses and responds to replication stress via ATR.
Transposition and cell cycle
Dr. Suneel Narayanavari*, Andrea Smith*
A temporary arrest at the G1/S transition phase of the cell-cycle was found to enhance transposition, suggesting that SB transposition is favored in the G1 phase of the cell-cycle, where the NHEJ pathway of DNA repair is preferentially active (Izsvak, 2004; Walisko, 2006). Recent findings indicate that, in addition to G1/S arrest, a temporal arrest in G2/M might also induce transposition. Notably, in contrast to certain viruses, severe DNA damage (e. g. DSBs) does not seem to trigger SB transposition, reflecting different strategies of various parasites that piggyback cellular processes.
Discovery of a novel phylogenetically-restricted cell type consistent with the human early embryo being a clonal selection arena
Manvendra Singh*, Aleksandra Kondrashkina, Jichang Wang*, Christine Römer*
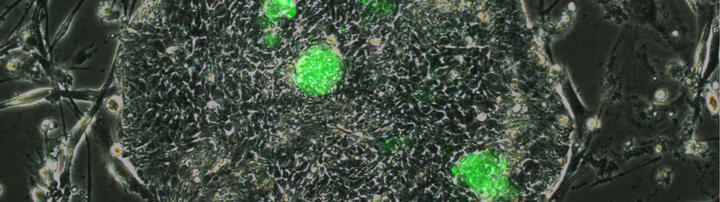
Clonal selection arenas, such as oocyte atresia, involve transposable element (TE) integration. Both apoptosis and transcripts from transpositionally-competent TEs occur in the early human embryo (EHE). We therefore ask whether there might be TE-mediated selection within the EHE. Single-cell analysis and embryo visualization uncover a common cell-type that segregates shortly after embryonic gene activation with the expected hallmarks. Prior to apoptosis these non-committed cells (NCCs), in contrast to their ontogenetic sisters, express transpositionally-competent Young TEs and DNA-damage response genes. Manipulation of a human embryonic cell line to mimic this novel type demonstrates TE transposition inducing DNA damage and pre-apoptosis. The viable ontogenetic sisters form the inner cell mass (ICM) and express suppressors of Young TEs including APOBEC and the non-transposing TE HERVH. NCC discovery enables improved characterization of ICM. With no similar evidence in mice, comparative early embryology provides a model to understand the phylogenetic occurrence of clonal arenas.
In collaboration with Laurence Hurst (Milner Center, Bath, UK and Jose Garcia-Perez University of Edinburgh,
bioRxiv 318329; doi: https://doi.org/10.1101/318329
Epigenetic regulation by human Endogenous Retrovirus H (HERVH)
Karam Ibrahim
Expressed transposable elements, including Endogenous Retrovirus-derived elements can fine-tune transcription by providing binding sites for transcription factors, but less is known about their role as independent regulators. Pre-implantational human embryos and cultured embryonic stem cells (hESCs) are expressing a plethora of long non-coding (lnc)RNAs, with only few of them being functionally characterised. In early human embryogenesis, Human Endogenous Retrovirus Family H (HERVH)-derived lncRNA are abundantly expressed, peaking in the epiblast cells and declining during differentiation. HERVH is also expressed at high levels in cultured hESCs and induced pluripotent stem cells (iPSCs). Depletion of HERVH expression results in differentiation and loss of self-renewal. Here, we aim to decipher the pattern of epigenetic regulation by HERVH via high-throughput Chromatin Isolation by RNA purification (CHIRP-seq). This approach would help us to identify the genomic regions targeted by specific HERVH-derived transcripts, and their role in regulating pluripotency.
Transposed Elements aid embryonic development
Manvendra Singh*
Transposable or Transposed elements (TrEs) are a productive source of biochemically active non-coding or occasionally coding elements that are tightly regulated in a cell-type specific manner. Many recent studies strengthen the hypothesis that few of these elements are co-opted for the regulation of host genes. We focus on one particular large family of human ERVs, which we find as a key regulatory player in the human pluripotent stem cells. I show that its co-option has potentially led the evolution and development of human-specific embryogenesis including the pluripotency. We updated transcriptomic encyclopedia of early human embryogenesis with their transcriptional flags. We re-define the progression of human embryogenesis at single-cell resolution with their markers. In conjecture with previous definitions, we characterize an unattended cell population in human preimplantation embryos that did or did not commit to any of the known lineages in order to form the stable blastocyst. Human transposcriptomics illustrate the contrasting pattern of transposon families during the progression of preimplantation embryogenesis. Their cross-talk with host factors could have driven the commitment of host cells to the particular lineages, leading to rapid turnover, compared with another species.
Functional significance of Sleeping Beauty transposase – HMGXB4 interaction
Guo Young*
In our previous studies, we identified HMG2l1/HMGXB4 as an interactor of the Sleeping Beauty transposase. Phylogenetic tree analysis shows HMGXB4 is a conserved gene from fish to human. In our follow up studies, we are trying to decipher the potential functions of HMGXB4, which may help us to understand the functional significance of such interaction between Sleeping Beauty transposase and its interacting Partner.
HMGXB4 Targets Sleeping Beauty Transposition to Vertebrate Germinal Stem Cells
Anantharam Devaraj* and Manvendra Singh*
We have shown earlier that HMGXB4/HMG2l1 (a component of the Wnt-signalling pathway) is able to physically associate with either the transposon DNA or the transposase protein, providing a negative feedback loop regulation to transposase expression (Walisko, 2008). New data suggest that HMXB4 has a high expression level in undifferentiated cells, but its expression level is dropping in differentiated cells in various models. We propose to validate the working hypothesis that HMGXB4 targets transposition to a well-defined developmental phase. The regulation of SB transposition in association with Wnt-signaling could be fundamentally different from LINE-1 (retrotransposon) regulation that is regulated primarily epigenetically in the human genome. The expression of HMGXB4 is conserved throughout the early development of human, rodents and flies. Which might explain the higher activity of TEs in the germ cells across the phylotree.
doi: https://doi.org/10.1101/2020.06.15.145656
Deciphering the function of HMGXB4 transcription factor in vertebrates
Guo Young
Phylogenetic tree analysis shows HMGXB4 is a conserved gene from fish to human. In our follow up studies, we are trying to decipher the cellular functions of HMGXB4.


