Ingo Morano
Molecular Muscle Physiology (Emeritus)
Profile
We perform that on the transcriptional, translational, and functional levels, using molecular biology, biochemical, cell-biological, and biophysical methods. Basic characterization of muscle functions of both sexes will help us to understand the distinct manifestations of muscle diseases.This is fundamental in order to develop new therapeutic strategies to treat muscle diseases, e.g. hypertrophic and dilative cardiomyopathies.
Therefore, our lab investigates the role of estrogen receptors (ER) on gene expression regulation in cardiomyocytes, in particular on the identification and functions of new co-regulators of the estrogen-activated ER, which modify the transcriptional activities of ER. To understand the energy balancing mechanisms of muscle, we are investigating the biogenesis, structure, and function of mitochondria in muscle.
Furthermore, we are studying the multitude functions of the key protein of Ca2+ activation of muscle, namely the voltage-dependent Ca2+ channel together with an interacting protein partner, the 700 kDa scaffolding protein Ahnak. The molecular mechanisms of depression of Ca2+ activation and, hence contractility of cardiomyocytes by the fatty acid binding proteins FABP3 and FABP4 are part of our ongoing research activities. Type II myosin and its associated light chains are the molecular motors of muscle contraction. Using transgenic animals we are investigating the role of normal essential myosin light chains (ELC) and mutated ELC associated with cardiomyopathy on cardiomyocyte contraction.
Contact
Research
Contracting adult rat cardiomyocyte electrically stimulated at 1Hz
Contraction of all muscle types is elicited by increasing myoplasmic Ca2+ and interaction of Type II myosins with thin (actin) filaments. In striated muscle, Ca2+ bind to troponin C, which turn the thin filament “on”, allowing actin-myosin interaction and force generation. The Morano lab is studying the functional roles of subunits of key proteins of Ca2+ handling and force generation, i.e. the L‑type Ca2+ channel, Ahnak, and type II myosins in muscle. Further, we investigating the effects and mechanisms of sex and the sexual hormone 17ß-estradiol in the heart in more detail. Particularly, we are screening new co-regulating proteins of estrogen receptor α and determine their effects in the heart. Any change of these proteins by mutation, differential gene expression, alternative splicing of the transcripts, or post-translational modification modulate muscle function. The ultimate goal of our research is to discover new effective strategies for the diagnosis, prevention, and treatment of cardiac diseases.
Myosin II is composed of two heavy chains (MYH) and four non-covalently linked light chains (MLC). The α‑helical lever arm of the MYH contains two IQ motifs in tandem. IQ1 binds the essential myosin light chain (ELC), whereas IQ2 binds the regulatory myosin light chain (RLC). ELCs consist of a lysine-rich N‑terminus, which binds to actin, and a C‑terminal lever arm binding domain which contains four helix-loop-helix EF-hand domains. In the adult human heart two ELC isoforms are expressed, namely an atrial-specific (MYL4, ALC‑1) and a ventricular-specific (MYL3, VLC‑1) isoform. Most patients with hypertrophic cardiomyopathy and congenital heart diseases re-express ALC‑1 in their ventricles, partially replacing the hVLC‑1 isoform which induced a pronounced positive inotropic effect. Hypertrophic cardiomyopathy (HCM) remains the most frequent genetic disorder of the myocardium and is the most common cause of sudden death in young people which manifests as a left ventricular hypertrophy without obvious cause. HCM associates with 12 missense mutations in MYL3. The functional consequences of MYL3 mutations as well as the pathomechanisms leading to HCM, however, are not clarified. Likewise, two MYL4 mutations which associate with atrial cardiomyopathy were reported recently with a similar lag in knowledge on the molecular pathomechanism.
Adipocyte fatty acid-binding protein (FABP4) is a member of a highly conserved family of cytosolic proteins that bind with high affinity to hydrophobic ligands, such as saturated and unsaturated long-chain fatty acids and eicosanoids. Recent evidence has supported a novel role for FABP4 in linking obesity with metabolic and cardiovascular disorders. In this context, we identified FABP4 as a main bioactive factor released from human adipose tissue that directly suppresses heart contraction in vitro. As FABP4 is known to be a transport protein, it cannot be excluded that lipid ligands are involved in the cardiodepressant effect as well, acting in an additional and/or synergistic way.
The cardiac voltage gated L‑type Ca2+ channel (CaV1.2) constitutes the main entrance gate for Ca2+ that triggers cardiac contraction. Several studies showed that the distal C‑terminus fragment of the α1C subunit of Cav1.2 (α1C-dCT) is proteolytically cleaved and shuttles between the plasma membrane and the nucleus, which is regulated both developmentally and by Ca2+. However, the effects of sex and sex hormone 17β-estradiol (E2) on α1C-dCT nuclear translocation are still unexplored. β‑Adrenergic stimulation enhances Ca2+ currents via CaV1.2 channels, strengthening cardiac contraction. The signalling via β‑adrenergic receptors involves elevation of cyclic AMP (cAMP) levels and activation of protein kinase A (PKA). However, how PKA affects the channel remains controversial. Recent studies in heterologous systems and genetically engineered mice stress the importance of the α1C-dCT.
Current Projects
- Characterization of new cardiodepressant factors from adipocytes (in cooperation with Dr. Valeria Lamounier-Zepter, TU Dresden)
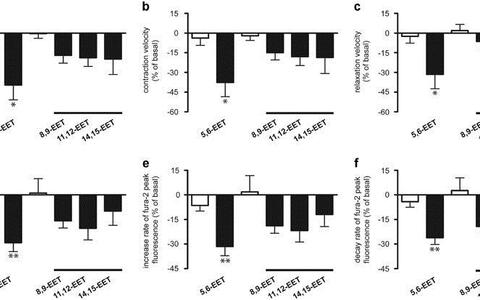
We investigated a possible involvement of lipid ligands in the negative inotropic effect of adipocyte factors in vitro. We verified that blocking the CYP epoxygenase pathway in adipocytes attenuates the inhibitory effect of adipocyte-conditioned medium (AM) on the contractility of isolated adult rat cardiomyocytes, thus suggesting the participation of epoxyeicosatrienoic acids (EETs) in the cardiodepressant activity. Analysis of adipocyte-conditioned medium (AM) for EETs revealed the presence of 5,6‑, 8,9‑, 11,12- and 14,15-EET, whereas 5,6‑EET represented about 45% of the total EET concentration in AM. Incubation of isolated cardiomyocytes with EETs in similar concentrations as found in AM showed that 5,6‑EET directly suppresses cardiomyocyte contractility. Furthermore, after addition of 5,6‑EET to FABP4, the negative inotropic effect of FABP4 was strongly potentiated in a concentration-dependent manner (Figure 2). These data suggest that adipocytes release 5,6‑EET and FABP4 into the extracellular medium and that the interaction of these factors modulates cardiac function. Therefore elevated levels of FABP4 and 5,6‑EET in obese patients may contribute to the development of heart dysfunction in these subjects.
Legend to Figure 2: Effect of control (open bars) and 5,6‑, 8,9‑, 11,12- and 14,15-EET (filled bars; 30 nM each) on fractional shortening (a), contraction (b) and relaxation velocity © of cardiomyocyte contraction, calcium transient (d) and on increase rate (e) and decay rate (f) of calcium transient of isolated cardiomyocytes. Values are expressed as the percentage change of basal value (mean±s.e.m.), n=6 – 11 experiments for each concentration, *P<0.05, **P<0.01 in comparison to the effect of the corresponding control (Lamounier-Zepter et al., J Obes (Lond). 2015;39:755 – 61).
© Morano Lab- Estrogen-activated estrogen receptor supports nuclear translocation of the C‑terminal domain of the cardiac L‑type Ca2+ channel (α1C-dCT)
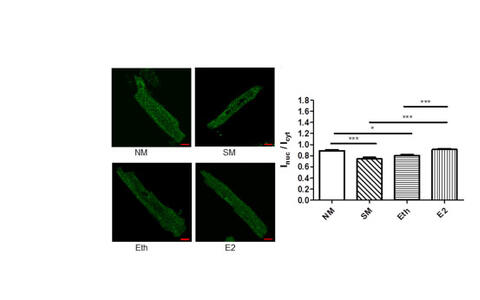
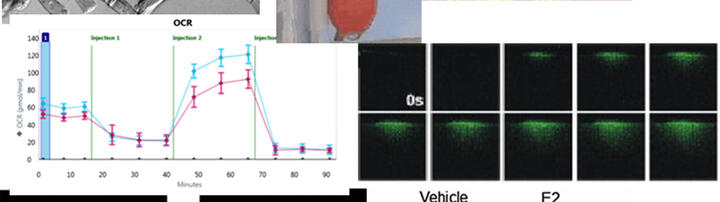
To investigate a possible sexual disparity in the α1C-dCT nuclear translocation, we first generated an antibody directed against a synthetic peptide (GRRASFHLE) located in α1C-dCT, and used it to probe ventricular myocytes from adult female and male mice. Immunocytochemistry of isolated mouse primary adult ventricular myocytes revealed both nuclear staining and cytosolic punctuate staining around the T‑tubules. The ratio of nuclear to cytosolic intensity (Inuc/Icyt) was significantly higher in isolated female cardiomyocytes compared to male cardiomyocytes. Western blot analysis of nuclear fraction confirmed these data. Furthermore, we found a significant decrease in nuclear staining intensity of α1C-dCT in both female and male cardiomyocytes upon serum withdrawal for 18h. Interestingly, subsequent E2 treatment normalized the intracellular distribution of α1C-dCT in male cardiomyocytes, but not in female cardiomyocytes. Treatment of male cardiomyocytes with E2 (45min) revealed a similar effect (Figure 3). This effect of E2 was revised by the estrogen receptor (ER) inhibitor ICI indicating the involvement of ER in this signaling pathway. Taken together, our results showed that the shuttling of α1C-CT in cardiomyocytes is regulated in a sex-dependent manner, and E2-activated ER may play a role in the nuclear shuttling of α1C-dCT in male cardiomyocytes. This may explain, at least partly, the observed sex differences in the regulation of cardiac Cav1.2 channel activity.
Legend to Figure 3: Left: Representative immunofluorescence images showing the effect of E2 on nuclear shuttling of α1C-dCT in male cardiomyocytes. Right: bars demonstrate the Inuc/Icyt ratios in male cardiomyocytes treated with M199 medium (with serum, NM), serum withdrawal (starvation medium: SM), vehicle control (ethanol, Eth) and 17β-estradiol (E2: 10−8 M) for 45 min. E2 treatment for 45 min led to a significant increase of α1C-dCT nuclear import. Values are means±SEM from at least 17 male cardiomyocytes per group. *p<0.05; ***p<0.001 (one-way ANOVA followed by Bonferroni’s test) (Mahmoodzadeh et al., J Mol Cell Cardiol. 2016, 97:226 – 34).
© Morano Lab- Characterization of ahnak1 actions on L‑type Ca2+ channel (in cooperation with Prof. Nathan Dascal, University of Tel Aviv, Israel).
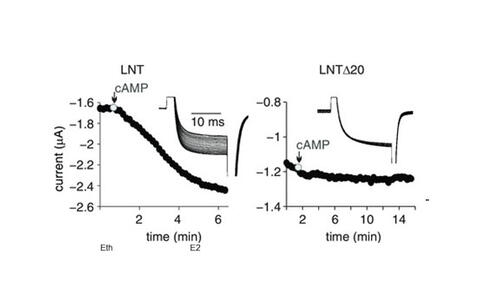
Here, we reconstituted the cAMP/PKA regulation of the dCT-truncated CaV1.2 in Xenopus oocytes. cAMP and the purified catalytic subunit of PKA, PKA-CS, injected into intact oocytes, significantly enhanced CaV1.2 currents. PKA blockers were used to confirm specificity. The regulation persisted in the absence of the clipped dCT (as a separate protein), the A kinase-anchoring protein AKAP15, and the phosphorylation sites S1700 and T1704, previously proposed to be essential for the PKA effect. The CaV β2b subunit was not involved, as suggested by extensive mutagenesis. Using deletion/chimeric mutagenesis, we have identified the initial segment of the cardiac long-N-terminal isoform of α1C (Figure 4) as a previously unrecognized essential element involved in PKA regulation.
Legend to Figure 4: representative time course charts of changes in IBa following cAMP injection in some of the constructs. Insets show current traces in the same cells. LNT denotes the long N‑terminal initial segment of α1C , LNTΔ20 a deletion mutant missing the first 20 N‑terminal aa of LNT (Oz et al. J Physiol. 2017, 595:3181 – 3202).
- The atrial essential myosin light chain modulates gene transcription by acting as a co-regulator of the E2/ERα complex
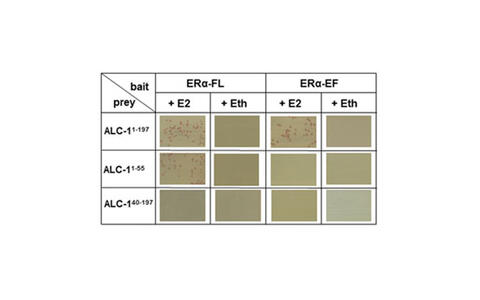
Chronic increased workload of the human heart causes ventricular hypertrophy and re-expression of the ALC‑1, an important positive inotropic factor of the human heart. Yeast two-hybrid screening of a human heart cDNA library revealed that E2/ERα complex interacts physically with ALC‑1 in the presence of 17β-estradiol (E2) (Figure 5). This interaction was confirmed by Co-Immunoprecipitation and immunofluorescence in human atrium. E2 induced the nuclear translocation of estrogen receptor alpha (ERα) and hALC‑1 in AC16 cells, where they cooperatively regulate the transcriptional activity of hALC‑1 gene promoter. E2-activated ERα required the estrogen response element (ERE) motif within the hALC‑1 gene promoter to reduce its transcriptional activity. This inhibitory effect was potentiated in the presence of hALC‑1 ~5 fold and thus, hALC‑1 acts as a co-repressor of ERα-mediated transcription. As a further novel effect, we showed that chronic E2-treatment of adult mouse cardiomyocytes overexpressing hALC‑1 resulted in reduced cell-shortening amplitude independent of Ca2+ activation levels. Together, our data showed that hALC‑1 acts as a co-regulator of the E2/ERα complex thus suppressing the transcriptional activity of the ALC‑1 gene.
Legend to Figure 5: Yeast two-hybrid analysis showed that only in the presence of E2 (10−8 M) full-length ERα (ERα-FL) interacts with the full length hALC‑1 protein (ALC-11 – 197) and the N‑terminus of hALC‑1 (ALC-11 – 55), while the EF domain of ERα interacts with the full-length hALC‑1 protein (1:1 retransformation assay) only in the presence of E2 (Duft et al., Basic Res Cardiol. 2017: 112)
© Morano Lab- Analysis of the gene expression profiles of cardiomyocytes regulated by the E2/ERα complex in the presence of human wild-type and mutated ALC‑1
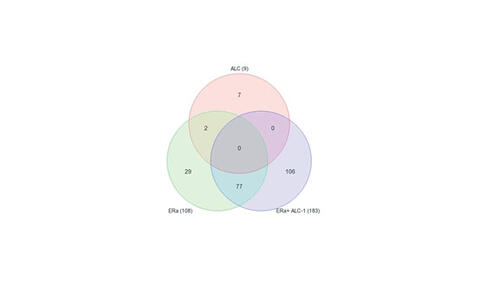
Since our recent data showed that ALC‑1 can also act as a co-regulator in the transcriptional machinery in the heart, we therefore aimed to profile the target genes, gene networks and pathways which are differentially regulated by E2-activated ERα/ALC‑1 complex in AC16 cells. The E2 effect analyzed in three transfection groups by paired two-way ANOVA revealed a list of 221 differentially expressed genes „DEGs“ (Figure 6). 137 genes were significantly differentially regulated in E2-treated cells either transfected with ERα or with ERα and ALC‑1. E2-ERα/ALC‑1 complex can either reverse the E2-ERα effects (31 genes) or regulates independently the gene expression in cardiomyocytes (106 genes). The aforementioned 137 DEGs were used for gene set analysis and pathway analysis (Partek® Genomics Suite® and Partek® Pathway™). Among others, one significantly enriched biological process (GO term) is “extracellular matrix organization”. One of the interesting pathways with a high enrichment score (4.4) is TGF‑β signaling pathway.
Legend to Figure 6: Venn diagram showing the number of differentially expressed genes when comparing E2-treated with vehicle-treated AC16 cells (limits: p<0.05; 2fold).
Disease-related mutations of MYL4, i.e. one frameshift and one missense mutation, have recently been described. Both MYL4 mutations associate with familial atrial cardiomyopathy with electrical, contractile, and structural abnormalities. We will therefore investigate whether the transcriptome of cardiomyocytes is differently regulated by those mutated ALC‑1 forms. The results will shed new light on the molecular pathomechanism causing the genesis of cardiomyopathy. We hope to identify disease-regulated genes which could explain the development of atrial cardiomyopathy caused by MYL4 mutations.© Morano Lab- Regulation of the secretory process of FABP3 of primary adult cardiomyocytes
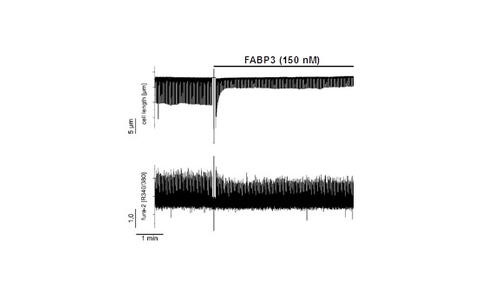
Fatty acid binding protein 3 (FABP3) is a cytosolic protein of cardiomyocytes, but also present in human plasma. Recent studies suggested a direct extracellular cardiodepressant effect of FABP3 on cardiomyocte contraction and Ca2+ handling (Figure 7). Interestingly, FABP3 has no secretory signal sequence; hence, the mechanisms how FABP3 is released from cardiomyocytes are unclear. In this study we will investigate the mechanisms of FABP3 secretion from mouse primary adult cardiomyoctes using specific blockers (brefeldin A, monensin A) of the Golgi-dependent secretory pathway in vitro, and raising intracellular Ca2+ levels (ionomycin). In addition, we will investigate the presence of FABP3 in exosomes and its releasing via this transport system from cardiomyocytes.
Legend to Figure 7: Inhibitory effect of FBBP3 on contraction amplitude (top) and intracellular calcium fluctuations (bottom) of a mouse cardiomyocyte.
© Morano Lab- Regulation of structure and function of mitochondria by Ahnak1 in the heart
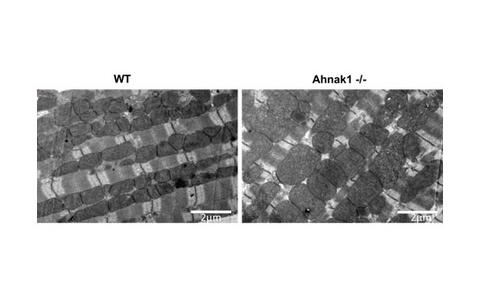
In the heart, Ahnak1 is located at the sarcolemma and T‑tubuli of cardiomyocytes indirectly associated with the voltage- dependent L‑type Ca2+ channel (L‑VDCC) via its β2-subunit (ß2), where it stimulates L‑VDCC current. Recent evidence has suggested that a high Ahank1 expression is associated with poor muscle fitness, although the underlying mechanism remains unclear. We investigate the role of Ahnak on physical fitness (4 weeks daily treadmill running training) of aged (12 months old) male and female mice using a homozygous Ahnak knock-out model (Ahnak-/-) and their Wild-type littermates (WT). Mitochondrial activity and phenotype of primary adult (aCM) cardiomyocytes of Ahnak-/- animals have been analyzed using the Seahorse technology and transmission electron microscopy (TEM). TEM study showed significantly lower mitochondrial number per area in female and male Ahnak-/- hearts in comparison to those of WT mice. However the Ahnak-/- hearts of both sexes have significantly more large (>1µm2) mitochondria and less small mitochondria (<0.5µm2) compared to age-and sex-matched WT hearts (Figure 8). Expression of OXPHOS proteins will be monitored by Western-Blot, expression of mitochondrial fusion proteins OPA1 and Mfn1/2, the fission proteins Drp1 and Fis1, and the key mitochondrial regulatory factors TFAM, PGC1 and Nrf1/2 by qPCR. Contractile parameters (maximal contraction amplitude, maximal rate of force development and maximal relaxation rate) and peak-systolic and diastolic Ca2+ levels of aCM prepared from Ahnak-/- will be compared with the corresponding WT groups. In addition, proteome analysis by Mass Spectrometry will be performed to identify those proteins regulated by ahnak1.
© Morano Lab