How cells move and don’t get stuck
Cell velocity, or how fast a cell moves, is known to depend on how sticky the surface is beneath it, but the precise mechanisms of this relationship have remained elusive for decades. Now, researchers from the Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) and Ludwig Maximilians Universität München (LMU) have figured out the precise mechanics and developed a mathematical model capturing the forces involved in cell movement. The findings, reported in the journal Proceedings of the National Academy of Sciences (PNAS), provide new insight for developmental biology and potential cancer treatment.
The mathematical model we developed can now be used by researchers to predict how different cells will behave on various substrates
Cell movement is a fundamental process, especially critical during development when cells differentiate into their target cell type and then move to the correct tissue. Cells also move to repair wounds, while cancer cells crawl to the nearest blood vessel to spread to other parts of the body.
“The mathematical model we developed can now be used by researchers to predict how different cells will behave on various substrates,” says Professor Martin Falcke, who heads MDC’s Mathematical Cell Physiology Lab and co-led the research. “Understanding these basic movements in precise detail could provide new targets to interrupt tumor metastasis.”
Teaming up to pin down
The finding comes thanks to experimental physicists at LMU teaming up with theoretical physicists at MDC. The experimentalists, led by Professor Joachim Rädler, tracked how quickly more than 15,000 cancer cells moved along narrow lanes on a sticky surface, where the stickiness alternated between low and high. This allowed them to observe what happens as the cell transitions between stickiness levels, which is more representative of the dynamic environment inside the body.
Then, Falcke and Behnam Amiri, co-first paper author and Ph.D. student in Falcke’s lab, used the large dataset to develop a mathematical equation that captures the elements shaping cell motility.
Previous mathematical models trying to explain cell migration and motility are very specific, they only work for one feature or cell type. What we tried to do here is keep it as simple and general as possible.
“Previous mathematical models trying to explain cell migration and motility are very specific, they only work for one feature or cell type,” Amiri says. “What we tried to do here is keep it as simple and general as possible.”
The approach worked even better than expected: the model matched the data gathered at LMU and held true for measurements about several other cell types taken over the past 30 years. “This is exciting,” Falcke says. “It’s rare that you find a theory explaining such a large spectrum of experimental results.”
Friction is key
When a cell moves, it pushes out its membrane in the direction of travel, expanding an internal network of actin filaments as it goes, and then peels off its back end. How fast this happens depends on adhesion bonds that form between the cell and the surface beneath it. When there are no bonds, the cell can hardly move because the actin network doesn’t have anything to push off against. The reason is friction: “When you are on ice skates you cannot push a car, only when there is enough friction between your shoes and the ground can you push a car,” Falcke says.
As the number of bonds increase, creating more friction, the cell can generate more force and move faster, until the point when it is so sticky, it becomes much harder to pull off the back end, slowing the cell down again.
Slow, but not stuck
The researchers investigated what happens when the front and rear ends of the cell experience different levels of stickiness. They were particularly curious to figure out what happens when it is stickier under the back end of the cell than the front, because that is when the cell could potentially get stuck, unable to generate enough force to pull off the back end.
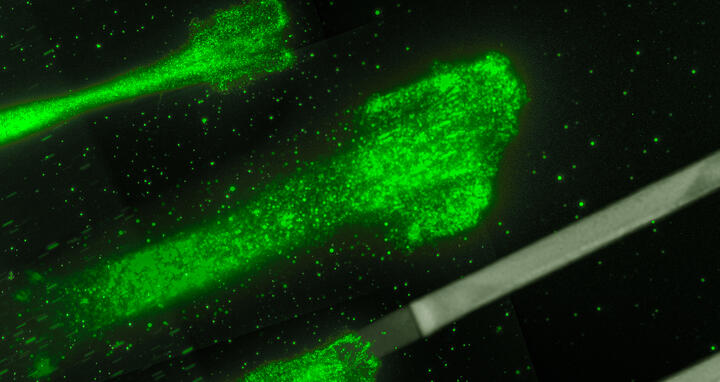
Cancer cells moving on glycoproteine strips: These strips act like splints, which allow to control and to study the movement of the cells better.
This might have been the case if the adhesion bonds were more like screws, holding the cell to the substrate. At first, Falcke and Amiri included this type of “elastic” force in their model, but the equation only worked with friction forces.
“For me, the most challenging part was to wrap my mind around this mechanism working only with friction forces,” Falcke says, because there is nothing for the cell to firmly latch onto. But it is the friction-like forces that allow the cell to keep moving, even when bonds are stronger in the back than the front, slowly peeling itself off like scotch tape. “Even if you pull just a little with a weak force, you are still able to peel the tape off – very slowly, but it comes off,” Falcke says. “This is how the cell keeps itself from getting stuck.”
The team is now investigating how cells move in two dimensions, including how they make hard right and left turns, and U‑turns.
Text: Laura Petersen
Further information
Literature
Christoph Schreiber, Behnam Amiri, Johannes C. J. Heyn, Joachim O. Rädler, and Martin Falcke: On the adhesion-velocity relation and length adaptation of motile cells on stepped Fibronectin lanes, DOI: 10.1073/pnas.2009959118
Download
Cancer cells moving on glycoproteine strips: These strips act like splints, which allow to control and to study the movement of the cells better. Rädler Lab, Ludwig Maximilians Universität München
Press contacts
Professor Dr. Martin Falcke
Head of Mathematical Cell Physiology Lab
Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC)
martin.falcke@mdc-berlin.de
Jana Ehrhardt-Joswig
Redakteurin, Abteilung Kommunikation
Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC)
jana.ehrhardt-joswig@mdc-berlin.de or presse@mdc-berlin.de
- The Max Delbrück Center for Molecular Medicine (MDC)
-
The Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) is one of the world’s leading biomedical research institutions. Max Delbrück, a Berlin native, was a Nobel laureate and one of the founders of molecular biology. At the MDC’s locations in Berlin-Buch and Mitte, researchers from some 60 countries analyze the human system – investigating the biological foundations of life from its most elementary building blocks to systems-wide mechanisms. By understanding what regulates or disrupts the dynamic equilibrium in a cell, an organ, or the entire body, we can prevent diseases, diagnose them earlier, and stop their progression with tailored therapies. Patients should benefit as soon as possible from basic research discoveries. The MDC therefore supports spin-off creation and participates in collaborative networks. It works in close partnership with Charité – Universitätsmedizin Berlin in the jointly run Experimental and Clinical Research Center (ECRC), the Berlin Institute of Health (BIH) at Charité, and the German Center for Cardiovascular Research (DZHK). Founded in 1992, the MDC today employs 1,600 people and is funded 90 percent by the German federal government and 10 percent by the State of Berlin.