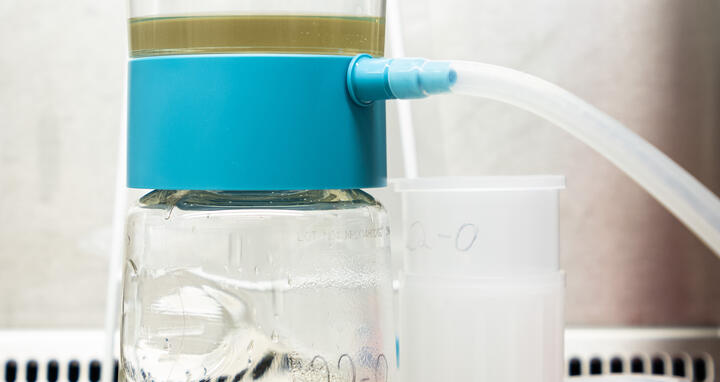📺 Covid updates from the sewer
Wastewater sample before sequencing.
Anyone infected with the coronavirus, regardless of whether or not they are symptomatic, excretes the virus’s genetic material – and not just via breath or saliva: The RNA of SARS-CoV‑2 can also be found in the stool of those infected. And this information is quickly carried via wastewater to sewage treatment plants.
In February 2021, several teams from the Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) began searching for the genetic material of the novel coronavirus in Berlin’s wastewater. They sequenced these data, interpreted them, and presented the results visually in descriptive graphs. Along with other colleagues, Vic-Fabienne Schumann and Dr. Rafael Cuadrat from Dr. Altuna Akalin’s Bioinformatics and Omics Data Science Platform have now published the outcome of their joint work. Dr. Akalin coordinated the project. With this publication the tool developed at the MDC is now available to other interested scientists.
Realistic incidence rates
It was Professor Nikolaus Rajewsky who came up with the idea of examining the murky waters of Berlin’s sewage system to obtain rapid and detailed information on the spread of the coronavirus in the German capital. The director of the Berlin Institute for Medical Systems Biology (BIMSB) at the MDC approached the Berliner Wasserbetriebe, which was only too happy to make its wastewater available for research purposes.

The resulting tool, a data analysis pipeline called PiGx SARS-CoV‑2, has now been described in detail by Akalin, senior author of the study, and his colleagues on the preprint server medRxiv. “This is a computer-based tool that allows us to graphically display infection dynamics and the SARS-CoV‑2 variants that are in circulation at different locations simultaneously,” Schumann explains. “The most important things to feed into this end-to-end pipeline are the results of the RNA sequencing from the wastewater, information about the relevant variants, and some ancillary information about the data.”
The results, which PiGx SARS-CoV‑2 displays as graphs, are not dependent on people taking Covid-19 tests or developing symptoms. They can also serve as an early warning system, Schumann explains, as they reliably predict whether incidence rates will increase or decrease over the coming days.
Early detection of new variants
To test their pipeline, the researchers analyzed a total of 38 wastewater samples from four Berlin sewage treatment plants between February and June 2021. “Using our tool, we were able to reconstruct the dynamics of the Alpha variant of concern, and we also detected the signature mutation of the Delta variant and its rise in early June,” Schumann reports. “We have therefore demonstrated that the pipeline works.”
Wastewater sequencing at the MDC is primarily carried out by Professor Markus Landthaler’s BIMSB lab RNA Biology and Posttranscriptional Regulation. “The major advantage with our computer-assisted methods is that we can simultaneously search for all known variants while also possibly detecting new mutations earlier than has been possible up to now,” explains Dr. Emanuel Wyler, a postdoctoral researcher in the Landthaler Lab. “Using the mathematical models we have developed, it may even be possible to detect variants of concern like Omicron before they become clinically relevant.”
Searching for Omicron
The MDC researchers will also be searching for the new Omicron variant. Schumann says they are currently in the process of sequencing the viral genome using samples from people infected with this variant. The team plans to use these results to refine the search and start tracking infection dynamics in early 2022. “There are still uncertainties surrounding how well our methods are able to identify entirely new variants,” explains the researcher. “It is not yet clear whether the viral RNA in wastewater is as complete as it is in the blood of patients.”
In any case, wastewater testing is not yet an established part of Germany’s Covid-19 early warning system – for either known or new variants. “Other countries like Sweden, the Netherlands and Italy are much further along with this,” says Schumann. “But perhaps our tool will help to change the situation in this country as well.”
Text: Anke Brodmerkel
Literature
Vic-Fabienne Schumann, Rafael Ricardo de Castro Cuadrat, Emanuel Wyler et al. (2021): „COVID-19 infection dynamics revealed by SARS-CoV‑2 wastewater sequencing analysis and deconvolution“. MedRxiv, DOI: 10.1101/2021.11.30.21266952
Please note: This manuscript is being made available to the scientific community on a preprint server. It has not yet been certified by peer review. As the authors may have to edit or make additions to the manuscript, it may take some time until journal publication occurs. Notwithstanding this, the MDC is presenting the research here as the subject matter is of high urgency due to the emergence of the Omicron variant of concern.