Two experiments in one fell swoop
Anyone who wants to know if two proteins bind together is spoiled for choice with nearly a dozen different methods to choose from. But one assay may be more suitable than others. This depends on such factors as the strength and duration of the interaction as well as on environmental conditions. “Unfortunately, each method can measure only a portion of all potential protein-protein interactions, ” says Professor Erich Wanker. “This is in each case only a small subset of the total number that occur.”
Together with his postdoc Philipp Trepte, the biochemist has now succeeded in merging several such subsets into a panorama analysis. Called LuTHy and described in the journal Molecular Systems Biology. The novel test procedure combines two established and complementary assays in one experiment. The combined method detects weak and strong interactions between proteins, as well as direct and indirect ones. Other methods reveal only whether a binding took place or not, while LuTHy also assesses the binding strength. What’s more, the findings of the two sub-experiments overlap – a feature that gives a better overall result.
Two proteins provide illumination
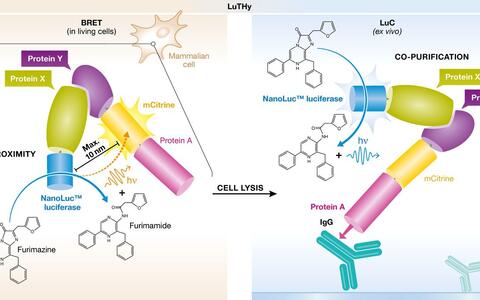
Like all two-hybrid methods, LuTHy employs two fusion proteins. It starts off by fusing fluorescent mCitrine and a protein A tag to the protein of interest. This part of the system is referred to as the “bait,” while the potential binding partner is called the “prey.” The latter is attached to nanoluciferase, an enzyme that generates light.
The nanoluciferase transfers energy to the fluorescent mCitrine only if the two proteins are in very close proximity, in which case mCitrine emits luminescence at a longer wavelength. The same phenomenon even occurs in living cells through a process called bioluminescence resonance energy transfer, or BRET for short. This part of the procedure measures weak or short-lived interactions that take place solely in the proteins’ natural environment.
The second part of the procedure yanks the proteins out of their natural environment. Only relatively strong interactions survive this process, in which the cells are disrupted and the fluorescent “bait” fusion protein is adsorbed on the wall of a microtiter plate well via the protein A. If the “prey” protein interacts with the bait, the luciferase it is carrying allows their association to be easily tracked. In interactions detected through luminescence‐based co‐precipitation, the fluorescent component mCitrine is no longer required and the distance between the two artificial light emitters is unimportant. Unlike with BRET, it is possible here to identify interactions in which the interaction partners are farther apart, such as those located within protein complexes or separated by adaptor proteins.
A method with potential
“LuTHy helps to address the commonly assumed weaknesses of the individual assays,” says the researcher Mikko Taipale in an accompanying commentary (“News & Views”). “It is quantitative, scalable, and compatible with small molecule treatments and diverse conditions, and it uses two independent measurements for detection of interactions.” Philipp Trepte is also pleased with the outcome: “We have created something along the lines of a new standard. This will make it easier to compare findings from different labs.”
What Wanker thinks is particularly interesting about this method is its application potential: “It allows us to examine how mutations affect interactions and disease development.” For example, he says, such a genetic change could convert a strong binding into a weak binding and do a great deal of harm to the cell. “Now when we test a small molecule with the assay and see that the interaction is strong again, that’ll mean we have a new drug target.”
Literature
Philipp Trepte et al. (2018): “LuTHy: a double‐readout bioluminescence‐based two‐hybrid technology for quantitative mapping of protein–protein interactions in mammalian cells.” Molecular Systems Biology 14, e8071. (Open Access) doi:10.15252/msb.20178071
Mikko Taipale (2018): “News & Views: Two protein/protein interaction assays in one go.” Molecular Systems Biology 14, e8485. (Open Access) doi:10.15252/msb.20188485