Structural basis of transcription regulation
Klf4 (Krueppel-like factor‑4) belongs to the family of SP/Klf zinc-finger transcriptions factors and is indispensable for terminal maturation of epithelial tissues. In a cocktail with transcription factors Oct4, Sox2 and c‑Myc, Klf4 can promote an opposite effect, the generation of induced pluripotent cells from differentiated tissues. The domain organization of Klf4 with amino-terminal proline-rich, acidic and PEST domains suggests that only the carboxy-proximal zinc-finger region adopts a compact, globular fold. A crystal structure analysis of the zinc-finger region bound to double-stranded cognate DNA reveals the basis of direct DNA sequence readout by Klf4. Each of the three classical CCHH zinc fingers contacts three base pairs of DNA in the major groove and achieves DNA sequence recognition by side chain-base hydrogen bonding (Figure). Six out of the ten base pairs contribute directly to sequence readout by Klf4, and the two carboxy-terminal zinc fingers are required for site specificity. It was shown that the lack of these two zinc fingers blocks the differentiation promoting activity of Klf4.
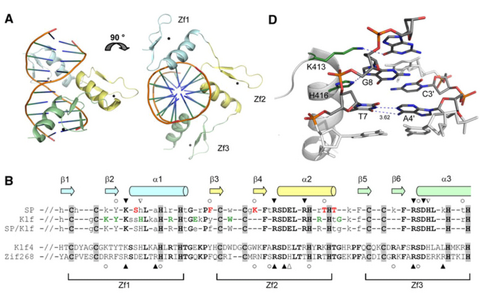
DNA sequence recognition by the zinc-finger domain of transcription factor Klf4. (A) Crystal structure of the Klf4 ZFD bound to double-stranded DNA with the consensus binding sequence. (B) Sequence alignment of the Klf4 zinc-finger region with the transcription factor Zif268 and consensus sequences for the SP, Klf and SP/Klf families. (D) Distortion of the DNA structure induced by the formation of a Klf4 His416·G8·C3’ pseudo-base triple.
Figure adapted from
Schuetz, A., Nana, D., Rose, C., Zocher, G., Milanovic, M., Koenigsmann, J., Blasig, R., Heinemann, U., Carstanjen, D.
The structure of the Klf4 DNA-binding domain links to self-renewal and macrophage differentiation
Cellular and Molecular Life Sciences (2011) 68, 3121 – 3131.