Alzheimer network
Protein-protein interaction network for Alzheimer’s disease
A characteristic feature of Alzheimer’s disease (AD) is the progressive neuronal cell death in the brain regions displaying high plasticity, caused by accumulation of intraneuronal neurofibrillary tangles (NFTs) and extracellular β‑amyloid plaques. NFTs consist of abnormally phosphorylated tau-protein, polymerized into paired helical filaments (PHFs), while amyloid plaques consist of the β‑amyloid peptide (Aβ1 – 42), which polymerizes into insoluble fibrils with high β‑sheet content. Modulators that influence tau- and Aβ-aggregation in vitro and in vivo are largely unknown.
The aim of this project is to create a protein-protein interaction (PPI) network for Alzheimer’s disease (AD). First, based on available literature information an interaction network will be generated using bioinformatic tools. Then, potential protein-protein interactions will be systematically tested using a membrane-based split ubiquitin system (Fig. 1). The split ubiquitin system identifies pair of interactions between hybrid proteins (baits and preys). Interactions are detected in yeast strains by activation of reporter genes. An automated version of the Y2H system will be applied to systematically identify protein-protein interactions.
The split ubiquitin system (modified from www.dualsystems.com)
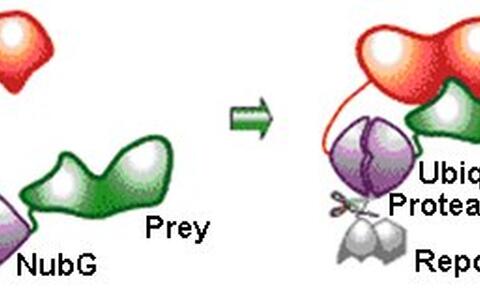
Fig. 1: The split ubiquitin system (modified from www.dualsystems.com)
The bait protein (in red) is fused to the Cub domain followed by a reporter protein, while a prey protein (in green) is fused to a mutated NubG domain. If bait and prey proteins interact, their interaction brings the NubG and Cub domains (in violet) into close vicinity allowing to reconstitute a ubiquitin protein complex. This allows the release of the reporter protein (grey) that functions as transcription activator.