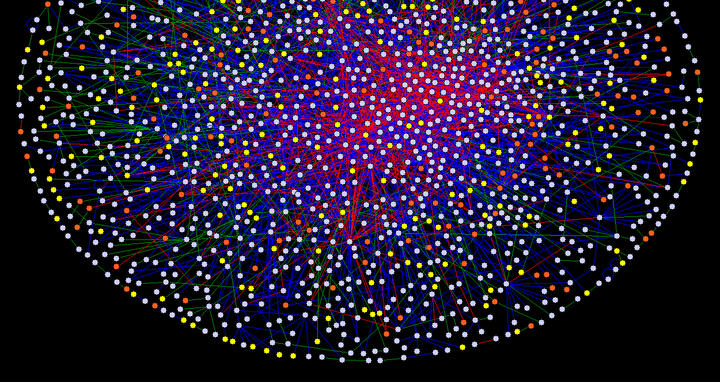
A human protein-protein interaction network: the first step towards a human interactome map
These macromolecules regulate the vast majority of cellular processes by their ability to communicate with each other and to assemble into larger functional units. Therefore, the systematic analysis of protein-protein interactions is fundamental for the understanding of protein function, cellular processes and, ultimately, the complexity of life. Moreover, interactome maps are particularly needed to link new proteins to disease pathways and the identification of novel drug targets.
Now, Ulrich Stelzl (laboratory of Prof. Erich Wanker) and colleagues have used an automated yeast two-hybrid system to screen more than 25 million pairs of human proteins for the identification of protein-protein interactions. The screen resulted in the creation of an interactome network consisting of 3,186 interactions among 1,705 proteins, representing a significant part of the human proteome. The map links 195 disease proteins to previously unidentified partners, allows the description of 342 uncharacterized human proteins via their interactions, and suggests new roles for hundreds of known proteins. New interaction partners (e.g., for the protein emerin that causes Emery-Dreifuss muscular dystrophy (EDMD) when mutated) hint at a novel function of the protein in membrane fusion processes.
In addition, the study integrates the interaction data with data concerning regulatory pathways, permitting extraction of potential functional complexes of proteins that participate in cellular signaling cascades. For example, the two proteins ANP32A and CRMP‑1, which are both implicated in human cancers, were linked to the Wnt signaling pathway. The paper published in Cell (Cell, DOI: 10.1016/j.cell.2005.08.029) is also a comprehensive resource for the functional characterization of human proteins and lays the foundation for a global connection scheme of the human organism.
Contact:
Pamela Cohen
p.cohen@mdc-berlin.de
+49 30 9406 2121