Learn to Fly
What others use as a breakfast condiment is available by the bucket load in Robert Zinzen's laboratory: Grafschafter sugar beet molasses. Add to this the dozens of liters of apple juice and the bags of white sugar, and you might think the man has a healthy appetite. But far from it: these ingredients are used to produce a very special diet. Fine-mesh netting in front of the door to an adjacent little room prevents Robert Zinzen's research objects from escaping (they are genetically modified). Ordinary fly catchers hang from the walls of the main lab to catch those that have managed to escape nonetheless.
Using the fruit fly model organism (Drosophila melanogaster), the researcher investigates how genes are regulated during the embryonic development of the nervous system. "A single fertilized egg cell develops into a complex organism with extremely differentiated cells and tissues that assume highly specialized tasks in spite of identical genetic material. How do cells decide what they are going to be over the course of embryonic development, and how do they get there?" the developmental biologist summarizes what is for him the central question. Since March 2013, he heads the junior research group "Systems biology of neural tissue differentiation" at the MDC's Berlin Institute for Medical Systems Biology (BIMSB).
Liver cell or nerve cell: the transcription factors decide
Amongst other factors, one group of proteins called transcription factors (TFs) is crucial for regulating whether a cell will develop into, for example, a liver or a nerve cell. Many TFs act by docking onto certain sections in the DNA and regulating the expression (i.e. transcription) of nearby genes. Combinatorially with other TFs, they control which information is expressed and since different cells contain different sets of TFs, they also feature different gene expression programs. Combinatorial regulation thereby allows hundreds of TFs to control the spatio-temporal transcription of tens of thousands of genes – a prerequisite for the proper development and existence of an organism. Fine-tuning of transcription, such as when exactly, where exactly, and how strongly a gene is expressed, is determined to a large extent via the exact interactions of TFs with the nearby DNA elements they bind to – the cis-regulatory elements (CRMs). Even small differences in TF availability can have major effects on a gene’s transcription, for example.
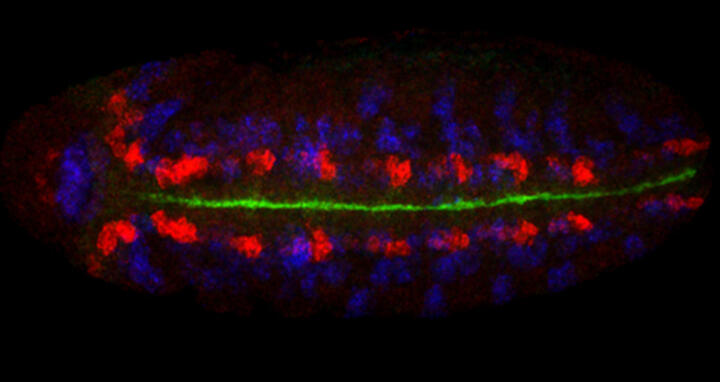
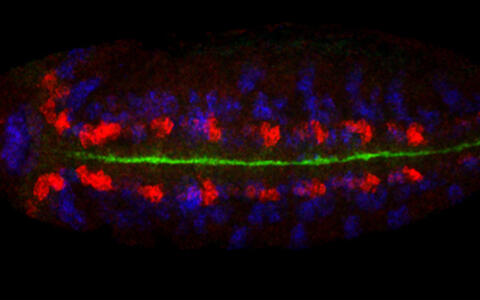
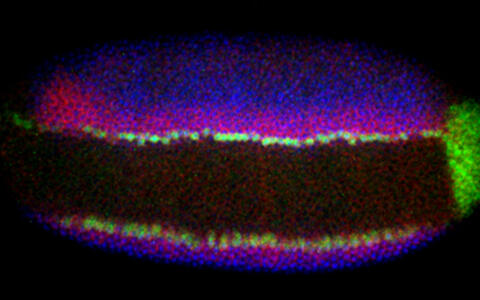
As soon as transcription starts in the fruit fly embryo, the process of cellular specification commences and within only a few hours a pool of pluripotent cells develops into differentiated components of the nervous system. The quick developmental progression as well as the excellent genetic and biochemical accessibility of the fruit fly embryo are some of the reasons why Robert considers these insects to be an ideal model organism for his research. "Many of the molecular processes we investigate are evolutionarily conserved from fly to man, which allows us to draw direct, testable inferences not only about nervous system development specifically, but also about tissue development in general" the scientist explains.
“Understanding how particular genes are expressed is really only a part of the puzzle”. The bigger challenge for the lab lies in deciphering the global networks and the interplay of TFs and CRMs across the entire genome: "So far, developmental biologists have often been limited to asking how individual genes are regulated; finally, though, we have the technological means to pursue the question of how the genome is regulated to direct and allow development." To this end, the researcher pursues an integrative systems biology approach: using the well-studied Drosophila model system, the lab produces diverse global datasets that are integrated using a multidisciplinary approach to formulate experimentally verifiable hypotheses about the regulatory networks. “One of the goals is to create an atlas of the regulatory interactions underlying nervous system development. Here, it is particularly important to find out how these regulatory structures change and adapt during the development process and how various types of cells subsequently differ from each other. The longer-term dream, however, would be the development a mathematical model of tissue development – one that not only explains the process, but one that would enable predictions of how certain disruptions and changes influence development and, for example, how synthetic gene constructs would behave," highlights the researcher.
Moreover, Robert Zinzen is in the process of establishing a fly facility at the MDC. This will enable groups without prior Drosophila experience to benefit from the know-how and equipment in his laboratory. Though this model organism combines many experimental advantages and is fairly cheap to maintain, the breeding and keeping of these insects can be somewhat labor-intensive. Embryos cannot be frozen, for example; rather, fly strains must be kept in small vials and transferred every month and their food must be specially prepared frequently. "Yet what makes Drosophila so attractive for me nonetheless – and this would also apply to other labs – is the sheer experimental accessibility, and the elegance and simplicity with which one can manipulate fruit flies. And don't underestimate the variety and wealth of Drosophila resources that have been established over a century of fly research. Having all this available in a developing and moderately complex organism is a dream-come-true for addressing many biological questions".
Cultivation of Drosophila has a long tradition on campus Buch
With Drosophila, Robert continues a long tradition of fly research on the campus in Buch. Already some 90 years ago, at the Kaiser Wilhelm Institute for Brain Research, radiation biologist and geneticist Nikolay Timofeev-Ressovsky used Drosophila as a model to investigate the nature of mutations and genetic variety. By way of applying X-rays he created mutations and examined the underlying genetic changes. In 1935, he and his colleagues Karl Günter Zimmer and Max Delbrück published the famous "Green Book", also called "Green Pamphlet", titled "Über die Natur der Genmutation und der Genstruktur".
Robert Zinzen knows he is in great company working at the MDC. "The MDC is a world-class facility and provides excellent conditions for scientists. In Germany, there are few comparable places, especially when it comes to systems biology" the 34-year-old researcher emphasizes. Born in the USA, Zinzen started his research in America. Last year he moved from Eileen Furlong's lab at the EMBL-Heidelberg to start his own lab at the BIMSB-MDC. He considers the interdisciplinary work on campus to be a great asset. "Good science rarely takes place in isolation, but rather is dependent on communication. Especially when it comes to our kind of questions, the colaboration with mathematicians, bioinformaticians and researchers from other fields is extremely productive; this is why the MDC generally, and BIMSB in particular was so very attractive."
Projects
With his team, Robert Zinzen works on a range of projects. Amongst others: Establishing a biochemical assay for isolating specific genomic sequences from developing embryos. Using transgenic flies, the aim is to isolate specific CRMs including bound TFs. "Here, it is very important that we achieve this in a tissue-specific manner. Since tissues don’t develop properly in isolation, but only within the embryonic context, it is necessary to understand developmental mechanisms in the embryo itself, rather than in culture," he says.
Another project concentrates on the creation of specific antibodies for neurogenic TFs that control the development of various embryonic nervous system precursors . Using ChIP-Seq (Chromatin ImmunoPrecipitation followed by Sequencing), the genome-wide binding profiles as well as their temporal binding dynamics and combinatorics are being investigated to better understand some of the basic aspects of how key TFs direct the underlying regulatory networks.
Another project deals with the modifications of histone proteins. Histones constitute protein complexes around which the genetic material can wind and condense. For a time, these structures were considered to be mainly packaging units. Today, we know that histone complexes are crucially involved in gene regulation, particularly through chemical modification of the histone tails. "Certain chemical modifications of these histone tails are highly indicative regarding the regulation status of genomic regions; genome-wide modification data can yield an entirely new level of information regarding genome regulation," the scientist emphasizes.
Featured Image: Not so different: developmentally flies and human beings have lots of common aspects. Shown here is a tripple-mutant Drosophila female with curly instead of straight wings, bar-shaped instead of round eyes and yellow body pigmentation. Photo: Robert Zinzen/ MDC