Zipping up toxic proteins
In hereditary diseases known as spinocerebellar ataxias, a protein called ataxin‑1 “zips up” into large, toxic clusters. Other molecules help it do so, and the labs of Erich Wanker and Miguel Andrade have now identified 21 other human proteins that can affect this process. They also found a common motif in the molecules that permits them to bind to ataxin‑1. The work, published in the August 16, 2012, edition of PLOS Genetics, provides important new insights into deadly neurodegenerative diseases and suggests new targets for combatting them.
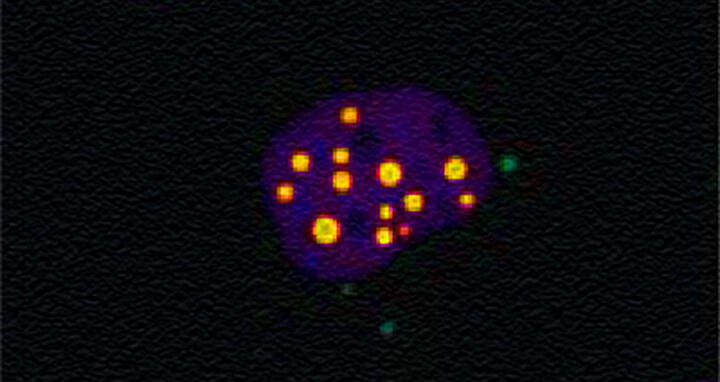
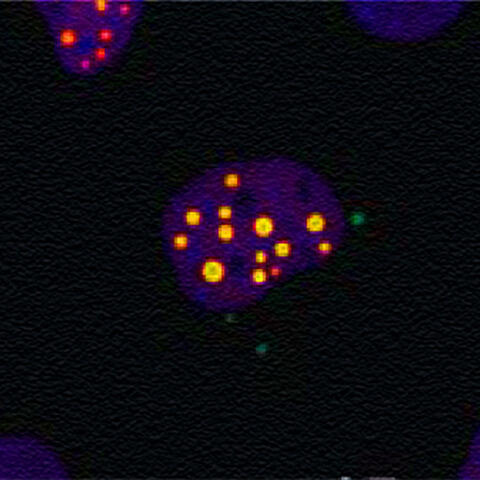
This microscope image revealed that the proteins ataxin‑1 and MED15 move to the same region of human cells, where they appear to interact and increase the way ataxin‑1 aggregates into toxic clusters.
Proteins are constructed as string-like chains of amino acids based on information in messenger RNA molecules and genes. The string quickly folds into a precise three-dimensional shape. This happens partly because of chemical attractions between the amino acids, partly through the help of other molecules. Problems with folding may give a protein the wrong shape, with an effect like twisting a piece of a puzzle. The misshapen protein may no longer fit the proper docking sites on other molecules; instead, it may latch onto other copies of itself and form clusters. Either situation can lead to disease.
Ataxin‑1 has a feature that is found in a few other human proteins and is closely tied to their toxic effects. Within the string, there is a long stretch that consists uniquely of a single amino acid called glutamine – as if you have held down a key on a computer and the same letter gets repeated over and over. Biologists represent glutamine with the letter Q so this segment is known as a polyQ sequence. It becomes dangerous if a person inherits a version of the molecule with over 40 glutamines in a row (called a polyQ-expanded version because it is longer than the healthy form). This feature causes it to fold in an abnormal way and zip up into clusters. The same problem affects huntingtin, which leads to Huntington’s disease in people who inherit a version with too many glutamines.
Why does the polyQ expansion change the way the protein folds and whether it forms clusters? This doesn’t happen on its own. Using laboratory organisms such as yeast, flies, and mice, scientists have discovered a number of other proteins whose behavior influences the process. Their effects on a mutant form of ataxin‑1 with an expanded polyQ may be positive or negative in terms of the protein’s clustering and toxicity. The labs of Miguel and Erich used computational methods to identify 192 close relatives of these proteins in humans, then added 45 human molecules that had been shown to bind to polyQ-expanded toxic proteins in protein-binding experiments.
This provided a list of 237 molecules that could potentially bind to the toxic form of ataxin‑1 and modify its aggregation and toxicity, but did they really do so in the cells of humans and mammals? Spyros Petrakis, Tamás Raskó, and Jenny Russ from Erich’s lab carried out experiments in cultures of mammalian cells and showed that 21 of these molecules modify the toxic effects of polyQ-expanded mutant version of ataxin‑1.
“The extra glutamine repeats probably create a special folded structure that permits the other proteins to dock on,” Erich says. “This suggested that those other proteins might have a common architecture that permits it, so our next step was to try to characterize it.”
The scientists approached the problem by comparing the architecture of molecules that enhanced the toxicity of mutant ataxin‑1 to that of molecules that decreased it. They were particularly interested in a structure called a coiled-coiled (CC) domain, a small region within many proteins that folds up to form two or more small helical “springs” lying alongside each other.
“Recent reports had suggested that these structures in proteins are critical for the accumulation of disease proteins with long glutamine repeats,” Erich says. “When we studied the molecules on our list, we found that half of those that increased toxicity likely had CC domains, whereas none of the molecules that decreased toxicity had them.”
These results were obtained through a computational analysis of the sequences of the molecules, in which Miguel and his colleagues discovered patterns of amino acids known to wrap into coils. Now they had to be followed up in the lab.
The scientists tested the effects of the CC domain in several ways. First, they found that one of the toxicity-enhancing molecules, called MED15, could cause the mutant ataxin‑1 to form clusters in the test tube. The same thing happened in cells. When they created an artificial version of MED15 without the CC domain, the effect disappeared.
The next step, Erich says, will be to determine exactly when and how MED15 and other molecules act in the cell to make toxic polyQ-expanded proteins more dangerous. “If we dissect the process and pinpoint these effects,” he says, “it will help us identify very specific regions of molecules and mechanisms whose behavior contributes to aggregation.” And that might provide precise new targets for a drug or therapy. Something that prevents specific CCs from binding to expanded polyQ sequences might inhibit the formation of clusters before they become toxic.
From the point of view of the collaborators, the project is already a success. Miguel says, “It shows that this strategy of combining computational analysis and experiments can help us identify additional molecules that are crucial in a disease process such as the disruptive aggregation of proteins.”
- Russ Hodge
Highlight Reference:
Petrakis S, Raskó T, Russ J, Friedrich RP, Stroedicke M, Riechers SP, Muehlenberg K, Möller A, Reinhardt A, Vinayagam A, Schaefer MH, Boutros M, Tricoire H, Andrade-Navarro MA, Wanker EE. Identification of human proteins that modify misfolding and proteotoxicity of pathogenic ataxin‑1. PLoS Genet. 2012;8(8)
Link to the free full text of the article