Thousands of new proteins found in dark proteome
Genes in DNA provide the recipe for cells to produce strings of amino acids, called peptides. Historically, peptides have been called proteins if they are long enough and have existing evidence for a biological role, such as the appearance of the same protein across species in evolution. A large, curated international database of proteins contains some 19,500 entities. But increasingly, scientists believe the traditional definition of a protein needs to be broadened.
A team of scientists led by the Princess Máxima Center for Pediatric Oncology, the University of Michigan Medical School, the EMBL European Bioinformatics Institute, the Institute for Systems Biology and collaborators at the Max Delbrück Center looked at more than 7,200 previously understudied sections of the DNA called non-canonical open reading frames (ncORFs). They found that some 25 percent of these sections – more than 1,700 – generated detectable protein-like molecules. These proteins are smaller than traditional proteins and therefore referred to as microproteins. The study was published in “Nature.”
Digging into the unknown
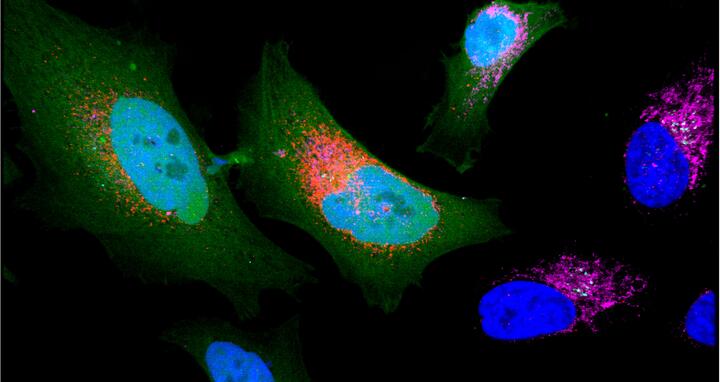
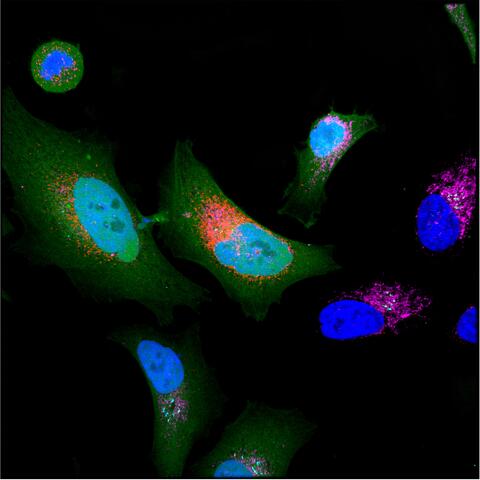
Cancer cell expressing microprotein (colored in red).
The scientists looked at 3.7 billion individual bits of raw data that may support known and previously unknown proteins – drawing upon 95,520 experiments, which took around 20,000 hours for computers to complete, working non-stop. They found 1,785 microproteins, a number that at first glance would increase the protein databases by nearly 10 percent.
But most of these 1,785 microproteins didn’t resemble the other 19,500 traditional proteins. For example, they were very small: 65 percent were fewer than 50 amino acids in length, compared to less than 1 percent of the 19,500.
Looking more closely at the microproteins, they saw that only a few – perhaps a dozen – resembled traditional proteins. For the remaining bulk, they spent over a year trying to figure out how to make sense of them.
The study builds on earlier research led by Dr. Jorge Ruiz-Orera, a senior postdoctoral researcher in the lab of Dr. Norbert Hübner, Group Leader of the Genetics and Genomics of Cardiovascular Research lab at the Max Delbrück Center. Hübner’s lab is a co-founder and a co-coordinator of the TransCODE Consortium, an international collaboration of more than 60 researchers from over 30 institutions.
Along with the lab of Dr. Uwe Ohler, Group Leader of the Computational and Regulatory Genomics lab, they identified more than 7,200 previously understudied ncORFs. The results, which were published in “Nature Biotechnology” in 2022, raised fundamental questions that led to the current study. Namely, how many of these regions produce stable protein-like molecules that are absent from the known human proteome? How many are biologically functional? And are they shared across many species in evolution, or are unique to humans?
Defining a new term
TransCODE consortium scientists have now coined a new biological concept: the peptidein. For decades, the research community has had a binary view of the relationship between human DNA and human proteins. A given piece of DNA either does or does not produce a protein. In their new study, the scientists propose a third option: DNA could make a protein, a peptidein, or neither.
The team defines a peptidein as existing in cells as a protein-like molecule, meaning that like proteins they are made of amino acids. But the role of a peptidein is ambiguous. Perhaps it has a function in normal human biology, or perhaps not; this is the key distinction from traditional proteins, where all are believed to have a function in normal human biology even if the details of that function are not fully known yet.
Importantly, this definition of peptidein leaves the door open for it to become a defined as a “protein” in the future – that is, if scientists gather more evidence on it. To start exploring this idea, the team searched for peptideins without which cells cannot survive. These so-called pan-essential peptideins may be important candidate drug targets in cancer and other diseases.
Peptideins as drug targets
Using large-scale CRISPR gene editing, the scientists found six peptideins that looked promising. For example, one of these was a peptidein produced from OLMALINC, a genetic sequence previously thought not to produce proteins. When the researchers switched this gene off, 85 percent of more than 485 cancer cell lines showed impaired survival. The researchers confirmed that this effect comes from the peptidein itself, not the RNA molecule it sits on, and found that it plays a role in cell division and DNA damage response.
Many of the newly detected peptideins are presented on cell surfaces for recognition by the immune system, making them potential targets for cancer immunotherapy. A number of such molecules presented to the immune system are already under development as drug targets, and there is growing interest from both academia and industry in exploiting this new class of cancer antigens. Peptideins could also shed light on genetic diseases that conventional gene analysis has been unable to explain, simply because genetic diagnostics were unaware that these molecules were encoded by the human genome.
Text: adapted from a press release issued by the Princess Máxima Center for Pediatric Oncology
Dr. Norbert Hübner, Group Leader of the Genetics and Genomics of Cardiovascular Research lab at the Max Delbrück Center, says:
“We are entering a particularly exciting phase in biology. The discovery of hundreds of peptideins gives visibility to a vast and previously overlooked layer of the genome and greatly expands the known proteome. Understanding their roles could transform how we study human disease, including cardiovascular disorders, and may reveal entirely new therapeutic opportunities.”
Dr. Sebastiaan van Heesch, research group leader at the Princess Máxima Center for pediatric oncology, who co-led the study, says:
“We know that the current overview of recognized proteins doesn’t capture the full picture. With this study, we show that thousands of overlooked genetic sequences contribute to the dark proteome by producing a new class of protein-like molecules, microproteins, that had been missed before now. But for most of them, we don’t yet know what they do.”
“It felt really special to discuss and decide what to do with this new class of molecules, as we had gathered enough early evidence to suspect that they might be widespread across cell types and tissues. By classifying these molecules of unknown functionality as peptideins, we’ve given them a formal place in reference databases so the wider community can study them.”
“With growing interest in industry and academia, peptideins are at the center of multiple drug development initiatives. Similarly, we see them increasingly turning up as important players in diseases, including childhood cancers. We hope to inspire a new wave of research into peptideins and to unlock new insights and drug targets across human biology, particularly for the development of cellular immunotherapies and cancer vaccines.”
Dr. John Prensner, pediatric neurooncologist at the University of Michigan Medical School, who co-led the study, says:
“We’re just beginning to see what this ‘dark proteome’ has to offer. It’s like the trailer to a movie. We see the outline of a game-changing view of human biology. We’re incredibly excited that the coming years will open new doors to help solve and treat human diseases such as cancer.”
Dr. Robert Moritz, Professor and Head of Proteomics at the Institute for Systems Biology, who co-led the study, says:
“Our collaborative work represents a culmination of decades of investment from federal funding agencies in building the computational and data infrastructure needed to interrogate the proteome at truly unprecedented scale at the Institute for Systems Biology.
By deploying our battle-hardened Trans Proteomic Pipeline across nearly 100,000 mass spectrometry experiments encompassing 3.7 billion spectra — derived from the world’s collective publicly available mass spectrometry data, with the results housed within PeptideAtlas at ISB for the scientific community to view and share — we were able to confirm, with high confidence, the existence of more than 1,700 of these newly identified peptideins that would otherwise have largely remained invisible to science.”
“What excites me most is not simply that these molecules exist, but what their existence implies.”
“Biology has long relied on a relatively small cast of well-characterized proteins to explain the regulatory logic of the cell, but peptideins suggest that beneath that familiar layer lies an entire untapped layer of molecular actors whose functional roles in gene regulation, signalling, and cytopersistence, many we are only beginning to imagine. Given their smaller size and the diversity of cellular contexts in which they appear, I believe peptideins may prove to be among the most versatile and consequential regulatory molecules we have yet encountered in human biology. This is not the end of a search — it is the opening of a vast and fertile new territory for the entire scientific community to explore and exploit, and I look forward to seeing what the broader scientific community uncovers as these molecules, and many more that are yet to be confirmed, are brought into the light.”
Further information
- Press release of the Princess Máxima Center for Pediatric Oncology
- Miniproteins appeared from nowhere
- Unchartered territory in the human genome
- Two MDC scientists receive ERC grants worth several million euros
- Hübner Lab
- Ohler Lab
Literature
Eric W. Deutsch, Leron W. Kok, Jonathan M. Mudge, et al. (2026) “Expanding the human proteome with microproteins and peptideins,” Nature DOI: doi.org/10.1038/s41586-026 – 10459‑x
- About the TransCODE Consortium
The study is the work of the TransCODE Consortium, an international collaboration of more than 60 researchers at over 30 institutions worldwide, co-led and coordinated by the Princess Máxima Center for Pediatric Oncology in the Netherlands, the University of Michigan Medical School, the EMBL European Bioinformatics Institute in Hinxton, the Institute for Systems Biology in Seattle, and the Max Delbrück Center. The study was supported by funders including the U.S. National Institutes of Health, the National Science Foundation, Oncode Accelerator (a Dutch National Growth Fund program) and the European Union’s Marie-Sklodowska-Curie program.