The search for the VRAC
Some important scientific questions resist a solution for years, until a new technology appears that seems to put an answer within reach. Success depends on creativity – finding a way to translate an old question into a form that can be handled by the new method – but also on timing. And a bit of luck never hurts.
After several years of work, Thomas Jentsch’s lab has shown that the VRAC channel is composed of proteins called LRRC8s. Humans have five forms of LRRC8 (A‑E). To function, the channel must consist of LRRC8A and one of the other four forms. Combining them in different ways allows cells to build versions of VRAC with different properties.
About four years ago Thomas Jentsch thought that the time had come for an all-out assault on a question that had teased biologists for decades. The campus had been acquiring instruments that might be the key to the solution. But the clock was ticking; other labs had the technology, too, and were interested in the same question: How do cells reduce their volume?
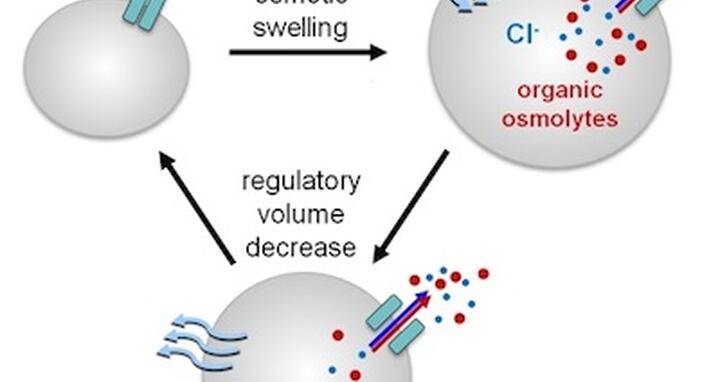
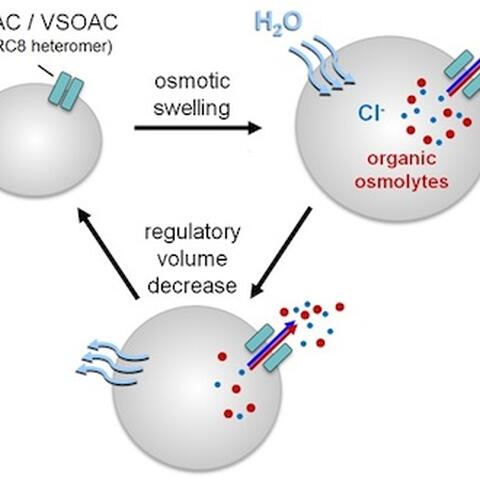
All types of cells undergo alternating phases of expansion and shrinkage as they divide, specialize, migrate, and cope with changes in their surroundings. In a liquid environment without any barriers, particles move from areas of high concentration to low. This applies also to water “particles”, which tend to diffuse until they have reached the same concentration everywhere. Water will flow from regions of high concentration (where it is diluted by a low concentration of osmolytes like ions, the constituents of salt, or other particles) to areas of low water concentration (where it is diluted by a high concentration of salt or other particles). So if cells are exposed to a hypoosmolar solution (which contains fewer osmolytes), water will flow into the cell, which now contains more osmolytes than its extracellular environment. But because the cell is enclosed by a membrane, the particles within it cannot simply be diluted to reach an equilibrium with extracellular concentrations. Instead, osmotic pressure will build up inside.
The same accumulation of internal water can occur when the number of internal particles increases through a breakdown of organic molecules, like glycogen to sugar, and cells may expand until the membrane is stretched to its limits. In order to keep from bursting, the cell needs to lose particles such as chloride or potassium ions, or organic osmolytes such as the amino acid taurine.
The loss of these particles decreases their internal concentration, which eventually becomes lower than in the external medium. This leads to a reversal of water flow, cell shrinkage and a recovery of the previous, normal cell volume and shape – called regulatory volume decrease. This volume decrease was known to be associated by a loss of negatively charged chloride ions (anions) from the cell through a channel.
Many aspects of the transport of chloride and other ions are well understood thanks to years of research on the part of Thomas’ lab and many others. This type of work has exposed hundreds of types of ion channels in cell membranes. These pore-like structures are composed of proteins woven through the membrane in a way that allows them to open and close. Channels are highly specialized to admit specific ions or other particles in response to changes in electrical charge, pressure, or other conditions. Their behavior is usually regulated by intricate biochemical networks. Most ion channels are so essential that any flaw in their components or their regulation can cause serious diseases – Thomas’ lab has established many such connections.
Much less was known about the outward passage of anions like chloride and other particles as cells shrink. Scientists postulated the existence of a specialized channel and had even given it a name: the volume-regulated anion channel, or VRAC. “VRAC was a hot topic,” Thomas says. “Hundreds of papers were being written about it and it was the subject of major international conferences.”
Despite all the interest, and the fact that the biophysical properties of the channel and some of its important functions were known, no one was able to actually find the channel protein. Occasionally a lab claimed to have identified a protein that belonged to it, but one by one, these candidates were discredited. While a number of drugs interfered with the channel’s behavior, they also disrupted other cellular processes. Without identifying a specific protein, scientists could understand neither these effects, nor the way swelling triggered changes in the channel’s behavior.
Part of the interest – aside from the fact that the channel served an important basic function in all types of cells – arose from observations that it, too, seemed to be linked to serious diseases. In any case, Thomas doubted that VRAC could hide much longer. The sequencing of the genomes of humans and other organisms had led to the development of technologies that permitted scientists to scan the DNA sequence for molecules with specific functions. Recent papers by other groups hinted that they were using these methods to search for the channel.
“Identifying the VRAC would finally give us a handle on some of these questions,” Thomas says. “Basically, it would open up an entirely new field.”
* * * * *
Until fairly recently, the study of gene functions almost always required an arduous, molecule-by-molecule approach. The earliest methods involved forward genetics: a scientist found an organism with a specific variant of a feature or a defect, then studied patterns of inheritance to establish that a single gene was involved. You might not identify it for years, or decades, but at least you knew it existed.
Modern biochemistry made it possible to do reverse genetics, through which scientists interfered with a specific gene and then studied the consequences for an organism. Most of these approaches depended on physically removing or altering genes, and thus eliminating the proteins they encoded.
The 1990s brought a powerful new alternative based on introducing artificial molecules called small interfering RNAs (siRNAs) into cells. siRNAs are composed of nucleotides, like other RNAs and DNA. This means that strands with complementary sequences will dock onto each other, which is the reason for the double-stranded helix structure of DNA. RNAs are usually single-stranded, but if an siRNA docks onto a complementary mRNA it produces a short, double-stranded region. Cells normally take notice and dismantle the molecule before it is used to make proteins. So artificially introducing siRNAs into cells has become a new, effective way of “knocking down” particular molecules by interrupting the flow of information from gene to RNA to protein.
Since the first applications of siRNAs in plants and simple model organisms, researchers have adapted them for use in human cells. The completion of the human genome revealed the entire set of mRNAs encoded in it, and scientists working in companies began constructing vast “libraries” of siRNAs to target them all. Theoretically this could be used to block the production of every human protein, one-by-one. Along the way, you would probably hit a molecule required for a particular function – such as the VRAC – and shut it down.
Humans have over 22,000 genes, meaning that the project would require at least that many experiments – actually the number would be much higher. There is no way to predict with certainty that a particular siRNA will work reasonably well within a cell. The way to get around this was to develop multiple siRNAs for each target and then challenge cells with two or three different versions; one of them would usually work.
But 22,000 experiments – let alone two or three times that number – could not be performed by hand. Robots and automation would be required at each step: to prepare samples, perform experiments and collect the data, and sophisticated software to analyze the results.
In the genome age, the technology and software needed for genome-wide siRNA screens began coming together, leading to several successful projects that have produced fascinating new insights into the functions of genes.
Several years ago the FMP created a Screening Unit, now jointly operated with the MDC, and acquired high-throughput, automated equipment to manage such large-scale projects. Headed by Jens von Kries, the facility has made significant contributions to a number of campus projects, many drawing on the tens of thousands of compounds that the unit has collected from researchers, pharmaceutical companies, and other sources. A typical use of this vast library is to challenge cells with substance after substance, hoping to disrupt a specific process in cells – with the aim of further developing the “hit” into a research tool, possibly even a drug. Chemists associated with the Unit then step in to tinker with a successful substance to strengthen its effects, make it less toxic for cells, or lend it other desirable properties.
The Screening Unit had also acquired a library of siRNAs that targets every known gene and then, through grants including a successful application from Thomas to the European Research Council (ERC), also a machine called FLIPR that was needed for following the time course of hundreds of experiments in parallel. So the necessary tools were on hand to search for VRAC components. “But first an extremely reliable experimental procedure had to be developed to detect the opening of the VRAC channel upon cell swelling,” Thomas says. “Any test that will be carried out tens or hundreds of thousands of times has to deliver clear, dependable results.”
Postdoc Tobias Stauber and PhD student Felizia Voss took the first steps toward developing a robust experimental protocol. This required almost two years, and as Thomas says, “There was no guarantee that we would find VRAC, even with this whole-genome approach, and certainly no guarantee that we would find it first.” Failure would have been particularly hard on Felizia, who was devoting her entire time to the project, and other PhD students who had become involved. Positive results would provide an impressive story for their dissertations; no one wanted to consider the alternative.
Tobias and Felizia began by growing cultures of human embryonic kidney cells – a favorite model for genetic research because the cells readily take up foreign molecules such as siRNAs. First they equipped cells with a yellow fluorescent “reporter” protein that would reveal whether cells changed their concentration of anions upon swelling. The cells were placed in a diluted medium in the presence of the ion iodide, which entered the cells through the opened VRAC channel and quenched the fluorescence. Although VRAC opening leads to a net loss of cellular chloride and volume regulation, the addition of external iodide, which is not present in the interior of the cell, leads to a net influx of iodide – channels are like “holes” that allow movement in both directions. The basic idea was that an siRNA that blocked the production of VRAC within the cell would abolish, or at least reduce, the loss of fluorescence upon exposure to a diluted medium containing iodide.
“This entire approach depended on an assumption that simply might not turn out to be correct,” Thomas says. “If a single specific molecule were required for VRAC, we might find it in the experiments. But if several molecules formed VRAC together, and each of them could be functionally replaced by another one, we might never find it with this approach. In fact, we discovered that VRAC has five components, and only one is absolutely required – we were lucky that only four of the subunits can replace each other.”
The scientists optimized the transfection of siRNAs and other aspects of the experiments, such as the number of cells, and established appropriate controls. Once this had been accomplished, it was time to scale up the experiment for a genome-wide search for VRAC components, trying to eliminate one essential component of the channel.
Now Katina Lazarow, a postdoc with the Screening Unit, and her assistant Sabrina Kleissle stepped in to adapt the procedure to the instrumentation. Each step of the pipeline of experimentation and analysis had to be adapted to the specific question the scientists were trying to answer. This required many months of intense work – often through the weekends. The actual screen would also take about two months of work, once again including weekends.
* * * * *
Such a large experiment required automation for the preparation of cell cultures, introducing a different siRNA into each one, then capturing images that would show whether the intervention had any effect on VRAC. These steps were performed in a specialized piece of machinery called the FLIPR, acquired second-hand with funds from Thomas’ grant and the Screening Unit.
The machine has become an important addition to the Screening Unit because it allows scientists to prepare experiments in all 384 “wells” of a assay plate in parallel, and follow changes in fluorescence over time, also in all the wells in parallel. The scientists needed to follow the exact time course of a decrease in fluorescence, because it is a measure of the flow of anions through VRAC.
Applied to the whole genome, this produced more than 130,000 fluorescence curves to analyze – a step that obviously required bioinformatic analysis. Miguel Andrade’s group at the MDC took on the task.
“We specified the parameters necessary to distinguish possible VRAC components from all the other proteins that didn’t affect the transport of chloride ions out of the cell,” Thomas says. “In the end we were left with a list of about 200 proteins that might be involved.”
More analysis and experiments were required to whittle the list down to just a few – hopefully just one – candidate. Part of this could be done by a computational analysis of the proteins’ sequences. The main goal was to find molecules with patterns indicating they were likely inserted into membranes. But other types of molecules might be worth investigating as well. A biochemical signal – passed along by a specific protein – might be required to operate the channel, even if that protein wasn’t a direct component of VRAC. Blocking it might shut down the channel just as well as eliminating one of the membrane proteins. The data collected in the experiments may eventually shed light on these processes. “We now have a treasure trove of data that remain to be analyzed; first we were concentrating on candidates for channel components,” Thomas says.
Felizia, Tobias and Thomas took on the list of candidate membrane proteins and began sorting through everything that was known about the functions of the candidates, from other experiments. The aim was to distinguish promising proteins from those that were less so – “And you had to be careful not to discard any molecule that might turn out to be the real one!” Thomas says.
After this analytical effort, over 80 proteins remained for one-by-one investigation through more experiments. Using the same experimental protocol, they challenged the cells with new siRNAs directed against those same 80 genes. The scientists dug in for another period of hard work.
This time, disruptions of only one molecule – a protein called LRRC8A – continued to interfere with VRAC. “So far we had only covered one aspect of VRAC function, the cells’ uptake of iodide,” Thomas says. “Now we wanted to explore another aspect – to measure its effects on electrical currents. We did this using a method called patch-clamp.” Two PhD students from Thomas’ lab, Florian Ullrich and Jonas Münch, joined the team and worked for about a year on the analysis of VRAC. They discovered that the knock-down of LRRC8A also reduced chloride currents. This meant that LRRC8A was either a direct component of the channel, or it exercised a firm control on VRAC functions. Thomas’ lab began the next round of experiments to find out.
* * * * *
“Proving that LRRC8A protein was a direct component of the channel required a number of steps,” Thomas says. “Upon a closer look we saw that it fulfilled a number of important criteria. First, it was expressed by many types of cells in the body, all of which need to regulate their volume and are thought to express VRAC. And a study of its sequence showed that it had regions that would be embedded in membranes – but was it really present in the plasma membrane that encloses cells?” The scientists generated antibodies that attached themselves to LRRC8A and observed under the microscope that LRRC8A indeed moved to its expected position at the outer membrane.
So far, so good – but was LRRC8A the channel itself? The scientists overproduced the protein in cells and measured currents activated by swelling. “Of course we had hoped that this would boost swelling-activated currents way beyond those in normal cells,” Florian says. “But were dismayed to see that rather than increasing currents, overproducing LRRC8A actually decreased currents!”
Thomas had an explanation for this behavior: “Channels are often composed of multiple proteins woven together in the membrane,” he says. “LRRC8A might form a channel together in a complex with other proteins, and if we had too much of LRRC8A, it may assemble with these other proteins at the wrong ratio, leading to complexes that cannot pass currents.”
What might those proteins be? Obvious candidates were molecules that were very closely related to LRRC8A. The cells of humans and other animals have many such “homologs”: far back in evolutionary history, in ancestral organisms, biochemical mistakes occasionally produced multiple copies of many genes. In the species in which they originally occurred, the copies would be redundant. That meant they could be lost again, which often happened as the genes naturally underwent mutations.
In some cases, however, spontaneous mutations enabled the copies to develop new functions. So the human genome contains four other versions of LRRC8 (labeled LRRC8B through E) that may function in a similar fashion.
“Databases containing information from other experiments showed that these molecules, too, are widely expressed in various types of cells,” Thomas says. “When we began to study these molecules individually, by introducing them into cultured cells, we noticed something interesting. Most of them remain inside the cell, in the cytoplasm. But if we introduced them along with LRRC8A, they move to the membrane.”
This indicated that LRRC8A binds to these related LRRC8 proteins, and that certain combinations of LRRC8s might be required to create the channel. Another round of experiments removed and restored various combinations of the proteins. Since siRNAs usually achieve only a partial blockage of a given protein, and might affect other molecules than the intended target, it was necessary to turn to another method. Here the scientists used a new technique called CRISPR-Cas, which interferes with genes directly at the level of DNA rather than its messenger RNA products. CRISPR-Cas allows scientists to make a physical cut in the genome and eliminate a specific gene. The results replicated the findings of the siRNA experiments and importantly showed other LRRC8s were involved in VRAC, but always in combination with LRRC8A.
The scientists found that each of the other LRRC8 proteins, B‑E, could be deleted alone without abolishing VRAC currents. But when B‑E were eliminated together, VRAC currents were gone just as if LRRC8A itself had been removed. “This showed that LRRC8A needs at least one of the other subunits to form a channel, but that LRRC8B through E are redundant – in other words, they can replace each other,” Felizia says. “And we even generated a cell line lacking all the forms A through E. Currents could only be restored when expressing A with at least one of the other family members.”
Importantly, different combinations of proteins yielded channels with different biophysical properties. “These are intrinsic channel properties,” Tobias says, “which demonstrates that LRRC8 proteins are not just involved in the signaling system that opens the channel upon cell swelling, but are an integral part of the channel itself.” This observation solved another enigma in the field – VRAC currents seemed to vary in different cells and tissues. “Our work now explains the variability of currents because different cells in the body make different amounts of these proteins. The different combinations may serve cell-specific functions.”
The project also provided hints toward answering another interesting question about channels involved in cell shrinkage. “The loss of volume involves expelling small organic molecules such as taurine,” Thomas says. “There was a controversy among experts – some claimed that these molecules exited via the VRAC, whereas others thought they might use yet another channel.”
Further experiments revealed that VRAC accomplishes both functions. The lab used to a radioactive form of taurine whose export from the cell could be tracked. Darius Lutter of Thomas’ group adapted a test that would measure its efflux, and the scientists found that taurine’s behavior aligned with the presence or absence of various LRRC8 molecules and VRAC behavior.
“All in all, these experiments reveal that the channel is directly constructed from combinations of LRRC8A and other LRRC8 forms,” Thomas says. “They give us the first clear route to explore the mechanisms by which cells shrink and shed particles in response to swelling. Those functions are crucial in the context of all types of cells, and this work provides the basis for understanding how they become defective in a number of serious diseases.”
Even with the well-coordinated collaboration and intensive efforts on the part of Thomas’ labs and the Screening Unit, the group’s search for the VRAC took four years. But the race would only officially be won if his team was first to publish on the subject. The scientists quickly wrote up a paper on the project and submitted it to a major journal.
All the way along, Thomas had been worried that other groups might be working on the problem; after four years, getting scooped would be a hard blow. Those concerns turned out to be justified. On the very same day that the article appeared in Science, another lab published a paper in the journal Cell that identified LRRC8A as a component of VRAC. That work did not, however, demonstrate that the “A” form had to be combined with another LRRC8 to create the channel.
“Our work puts the whole field of cell volume regulation on firm ground,” Thomas says. “The signals that connect an increase of cell volume to the opening of the VRAC are enigmatic, and we now have a handle to investigate them. We have suspected that defects in VRAC are involved in a number of diseases, and now we can develop mouse models to confirm those roles and look for other functions of the channel.”
– Russ Hodge
Highlight Reference:
Voss FK, Ullrich F, Münch J, Lazarow K, Lutter D, Mah N, Andrade-Navarro MA, von Kries JP, Stauber T, Jentsch. Identification of LRRC8 heteromers as an essential component of the volume-regulated anion channel VRAC. TJ.Science. 2014 May 9;344(6184):634 – 8. doi: 10.1126/science.1252826. Epub 2014 Apr 10.
Link to the full text of the paper