Comprehensive new map helps understanding of gene regulation by microRNAs
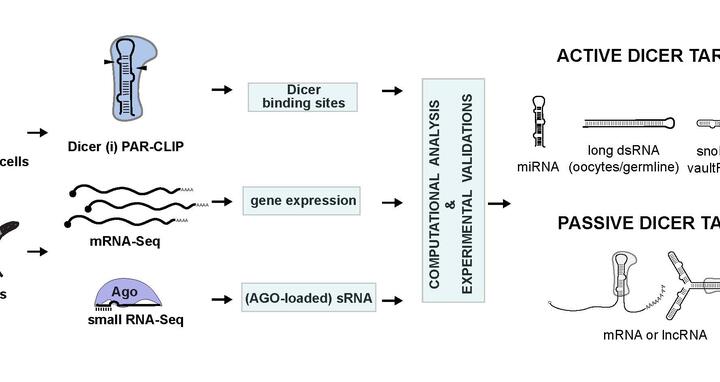
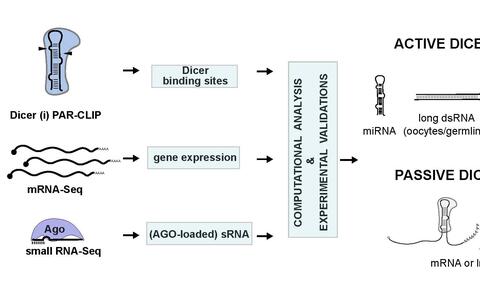
All the cells in our bodies, be they nerve or kidney cells, blood or skin cells, carry the same genetic information. And yet they all differ considerably. This is because different genes are transcribed in each cell type. In recent years, scientists have found a new class of molecules, so-called “microRNAs,” that play a crucial role in this gene regulation. An enzyme known as Dicer is involved in the formation of these microRNAs. A team of researchers led by Markus Landthaler and Nikolaus Rajewsky at the MDC’s Berlin Institute for Molecular Systems Biology (BIMSB) have recently compiled a comprehensive map of the RNA-binding and processing sites of Dicer, which they published in the journal Cell. Increased understanding of the basic mechanisms governing gene regulation could help in the search for targeted drugs and therapies.
When genes are read in the cells of our body, a transcript in form of a messenger RNA, a ribonucleic acid, is generated. This then serves as an instruction for the production of proteins. Recently, geneticists discovered another important class of small ribonucleic acids in cells, microRNAs. These molecules can bind to very short regions of the much longer messenger RNAs and thereby block them. Thus, microRNAs influence whether or not a protein is produced, and how much is produced. In previous studies, a team of researchers led by Nikolaus Rajewsky at the MDC demonstrated that, in this way, microRNAs regulate thousands of genes in our cells.
This knowledge about microRNAs is important because mistakes in gene regulation are instrumental in the development of many diseases. That is why we need to understand how these small molecules block the messenger RNA, thus turning off genes, and how microRNAs are created in the first place. For some time now, an enzyme known as Dicer has been identified as being involved in the process. Dicer cuts the actual microRNAs from so-called “pre-microRNAs.” These then form a complex with an Argonaute protein, which can dock onto messenger RNA and thereby block it. Researchers have discovered most of the 1,800 microRNAs identified so far by looking for RNAs bound to an Argonaute protein.
“We asked ourselves which RNAs Dicer actually binds to, which RNAs it processes, and whether there are other Dicer substrates besides pre-microRNAs,” says Markus Landthaler of the MDC. Landthaler’s team has not only found new microRNAs; working with Nikolaus Rajewsky’s group, they also succeeded in describing other previously unknown regulatory functions for Dicer. Landthaler and Rajewsky published their results in Cell last November.
The team led by the two scientists studied cells from the human kidney as well as cells from the nematode Caenorhabditis elegans. The latter is regarded as an important model organism because it allows for easy study of early developmental processes. Also, microRNAs were first characterized in the nematode. Researchers used biochemical methods to comb through the “transcriptome” of these cells – i.e. all the RNAs that are present at a given time. They found thousands of Dicer-binding sites, hundreds of which were located on known pre-microRNAs, and many more on pre-microRNAs that had not been characterized before. This way, they were able to identify more than 360 new microRNAs.
“What is interesting in this context is that we found several Dicer-binding sites in unexpected locations, namely on messenger RNAs,” says Landthaler. Apparently, Dicer processes some microRNAs that are not from pre-microRNAs, but from messenger RNAs.
Surprisingly, the researchers found a variety of Dicer binding sites on RNA regions that are not processed into microRNAs and therefore do not form a complex with an Argonaute protein. This discovery indicated that Dicer must have other functions. Indeed, the team led by Landthaler and Rajewsky found that Dicer apparently stabilizes some messenger RNAs and likely plays a role in localizing them. As these functions were observed in human cells as well as in C. elegans, the functions are evolutionarily conserved and are thus probably very important. The researchers also found a long, non-coding RNA in C. elegans that sequesters Dicer and thus influences the processing of microRNAs.
This research into the basic mechanisms of gene regulation could help find targets for therapies and drugs in the future. First, we have to understand the regulatory and additional functions of Dicer. The findings of Landthaler and Rajewsky and their team have made an important contribution to achieving precisely that.
- Elke Binder
Figure: Markus Landthaler and Nikolaus Rajewsky have recently compiled a comprehensive map of the RNA-binding and processing sites of Dicer. (Image: BIMSB)
Highlight Reference:
Rybak-Wolf, A., Jens, M., Murakawa, Y., Herzog, M., Landthaler, M., & Rajewsky, N. A variety of dicer substrates in human and C. elegans. Cell (2014) 159:1153 – 67. doi: 10.1016/j.cell.2014.10.040.