Project brings voluntary help into the lab
What is LabHive all about?
Dr. Tobias Opialla arbeitet am MDC in der “Proteomics and Metabolomics Platform” von Dr. Stefan Kempa am BIMSB.
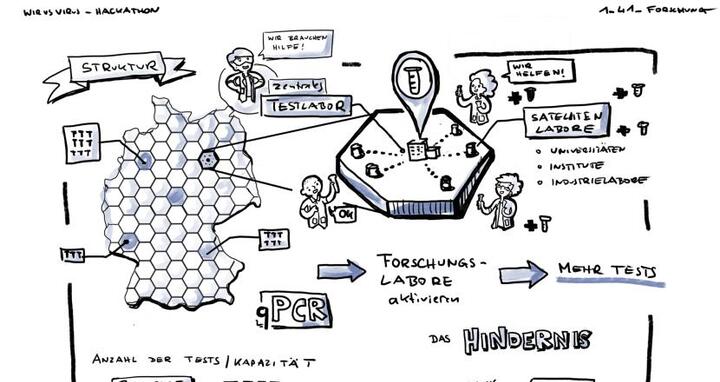
There are currently many scientists who cannot work in the laboratory at all or only to a limited extent. As a result, many labs are full of equipment that is not currently in use. We thought: This can’t be. On LabHive, qualified volunteers and labs can register for free and offer resources – be it labor, equipment, reagents, or experience with PCR and evaluating data. Diagnostic laboratories can then search these offers within their locality and using filter criteria. We know these diagnostic laboratories are still working close to their limits and that, even though diagnostic capacities have recently improved, there is still a need for support and advice. For example, no standardized national protocol currently exists for PCR tests. We want to help channel resources and needs.
How did the idea for the project come about?
A few weeks ago, an appeal for lab technicians was sent out from Labor Berlin – the largest diagnostic laboratory for Berlin hospitals. Within a few hours, several hundred volunteers had responded. Diagnostic laboratories calling for help have been flooded with offers. They are simply not able to answer each one individually, and it’s not nice for volunteers to receive no reply. I wondered whether it might be possible to process these requests and inquiries via a central platform, and signed up to participate in the German government’s hackathon #WirvsVirus. There, I was one of more than 28,000 volunteers working together to find solutions to help us through this crisis. I met many other people who also had the idea of pooling volunteer resources. We had 48 hours to develop a concept. From the 1,500 solutions submitted, our team qualified for the Enabler Program. This program is funding the implementation of 130 open source projects that were born out of the hackathon. Since then, we have also obtained funding from the BMBF.
How is it possible to get a project like this up and running in such a short time – especially during a global health crisis?
An interdisciplinary team coordinates test capacities for SARS-CoV‑2 using the LabHive digital platform.
We are now in the fifth week since the hackathon, which took place in the end of March. Everyone is doing this alongside their regular jobs. We are lucky to have many professionals on board – currently there are 15 of us from different companies and institutes from all over Germany. This includes, for example, a doctoral student who is familiar with virological issues, and a biotechnological assistant who knows about the needs of diagnostic laboratories. We also have people from the fields of data protection law and IT security on the team. Developing an IT security concept usually takes weeks. But when our partner, the Björn Steiger Foundation, recently requested it, we had it ready within 24 hours. It is rare for a hackathon project to meet high security standards right from the start. Moreover, the programmers, user interface designers and graphic designers on our team do this kind of work day in, day out. This helps everything run very professionally and we enjoy a good working relationship with the Björn Steiger Foundation, who is now responsible for bringing LabHive live.
By the way, we have never met as a team or even seen each other’s faces, as we always talk via group calls rather than video chats. This works incredibly well. You can still feel how motivated everybody is. In mid-April we also further developed the platform at #EUvsVirus and are getting a lot of feedback, also from abroad. It is incredibly rewarding to be working together with the common goal of helping to overcome this crisis.
Further information
- The LabHive platform
- Solution Enabler Program of the German government’s #WirVsVirus hackathon
- #EUvsVirus Challenge – a pan-European hackathon organized by the European Commission
- Tobias Opialla works at BIMSB: