Protein Production & Characterization
Dr. Anja Schütz
Profile
Our mission is to enable technology development and cutting-edge research by fostering collaboration with other technology platforms and research groups. The PPC Platform offers customized protein production for various downstream applications such as:
- Interaction studies
- High-throughput screening campaigns
- Tool protein production
- Antigen/Antibody production
- Structural biology projects
- Biochemical & biophysical characterization
Next to customized protein production, the PPC Platform is an ideal partner to set up collaborative research projects on topics such as protein engineering and protein characterization. We have a strong interest in studying the structure-function relationship of proteins linked to human diseases. By applying various biochemical and biophysical methods, we delineate the physicochemical properties of the analyzed proteins, a prerequisite to understand their biological function.
The PPC Platform is in close contact with other protein productions sites and partakes in international benchmarking studies initiated through the P4EU network.
Besides being an established partner for educational training of MDC-internal apprentices, the PPC Platform is recruiting bachelor and master students. In the future, we want to promote the success of interdisciplinary PhD projects by offering the possibility of co-supervision of doctoral theses.
Are you interested in a collaboration with us? We are looking forward to your enquiry!
Team
Alumni
We thank our former team members for their active support.
Berger, Ingrid – Technical assistant
Donath, Mandy – Bachelor student
Haake, Fabian – Bachelor student
Lenski, Ulf – Bioinformatician
Kabuß, Loreen-Claudine H. – MDC apprentice
Kuhnke, Alina – MDC apprentice
Kurths, Silke – Technical assistant
Mustroph, Mandy – MDC apprentice
Radusheva, Veselina – Master student
Rossa, Denise – MDC apprentice
Simon, Carolin – Master student
Service & Technology
We have established Standard Operating Procedures (SOPs) and attach particular importance to quality control. Our philosophy is to closely cooperate and actively exchange ideas with our collaborators, since this is essential for successful cooperation.
- Services
The following services are offered by the PPC platform:
Consulting
Scientific & technical level Molecular biology services
(1) Construct design
(2) Gene cloning
(3) Site-directed mutagenesis
Protein production services
(1) Small-scale expression & solubility screening
(2) Scale-up expression
(3) Protein purification
Protein characterization services
(1) Protein stability determination & optimization
(2) Protein-ligand interaction studies (ligand = protein, nucleic acid, small molecule)
(3) Oligomerization analysis
(4) Protein intact mass analysis of peptides and proteins via LC/MS TOF
(5) Protein structure determination (including crystallization, X‑ray data collection)
(6) Folding analysis
- Methods & Technologies
Protein Production
Gene expression
construct screening & large-scale expression
bacteria – E. coli (up to 48 L)
mammalian cells – HEK293, CHO (suspension, transient)
Protein purification
ÄKTA-FPLC chromatography systems
AC, SEC, IEX, HIC columns & resins
refolding
Protein Characterization
©Agilent Technologies, Inc. Reproduced with Permission, Courtesy of Agilent Technologies, Inc.
TOF LC/MS (Agilent)
intact protein mass analysis
©Wyatt Technology Corporation, Santa Barbara, CA, USA
DynaPro Plate Reader III (Wyatt)
protein stability determination & optimization (DLS/SLS)
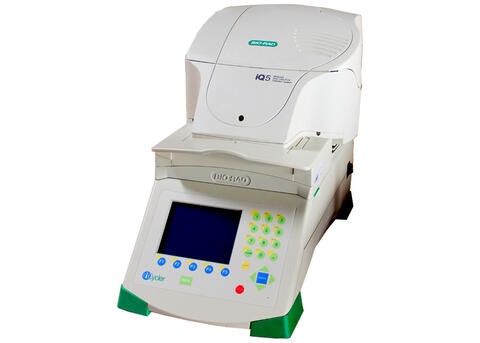
iQ5 RT-PCR (BioRad)
thermal shift assay (DSC)
MicroCal PEAQ-ITC (Malvern)
protein-ligand interaction studies
Chirascan CD Spectrometer pliedPhotophysics)
protein folding analysis
Protein Structure Determination
©ARI — Art Robbins Instruments, Crystal Gryphon LCP
Crystal Gryphon LCP
(Art Robbins Instr.)
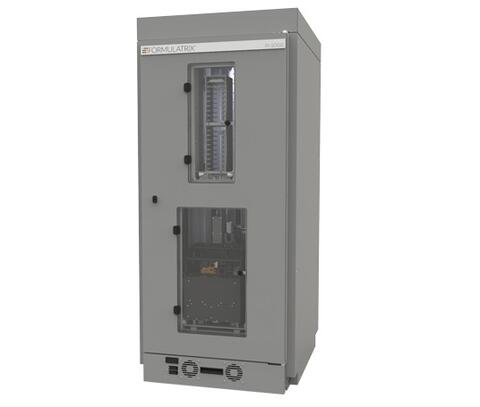
©FORMULATRIX® — Laboratory Automation Solutions Rock Imager 1000
Rock Imager
4°C und 20°C
(Formulatrix)
Access to synchrotron
(HZB — Bessy)
In case you are interested in protein-related services not explicitly listed here, please contact us.
Publications
- Peer-reviewed publications
- Acknowledgements
- Gustafsson O, Krishna S, Borate S, Ghaeidamini M, Liang X, Saher O, Cuellar R, Birdsong BK, Roudi S, Estupiñán HY, Alici E, Smith CIE, Esbjörner EK, Spuler S, de Jong OG, Escobar H, Nordin JZ, Andaloussi SEL. Advanced peptide nanoparticles enable robust and efficient delivery of gene editors across cell types. J Control Release. 2025 Oct 10;386:114038. doi: 10.1016/j.jconrel.2025.114038. Epub 2025 Jul 18. PMID: 40684990. doi: https://doi.org/10.1101/2024.11.27.624305Irastorza-Azcarate I, Kukalev A, Kempfer R, Thieme CJ, Mastrobuoni G, Markowski J, Loof G, Sparks TM, Brookes E, Natarajan KN, Sauer S, Fisher AG, Nicodemi M, Ren B, Schwarz RF, Kempa S, Pombo A. Extensive folding variability between homologous chromosomes in mammalian cells. Mol Syst Biol. 2025 Jul;21(7):735 – 775. doi: 10.1038/s44320-025 – 00107‑3. Epub 2025 May 6. PMID: 40329044; PMCID: PMC12222873.Extensive folding variability between homologous chromosomes in mammalian cells — PubMedWaltho A, Popp O, Lenz C, Pluska L, Lambert M, Dötsch V, Mertins P, Sommer T. K48- and K63-linked ubiquitin chain interactome reveals branch- and length-specific ubiquitin interactors. Life Sci Alliance. 2024 May 21;7(8):e202402740. doi: 10.26508/lsa.202402740. PMID: 38803224; PMCID: PMC11109483.K48- and K63-linked ubiquitin chain interactome reveals branch- and length-specific ubiquitin interactors — PubMedLebedin M, Ratswohl C, Garg A, Schips M, García CV, Spatt L, Thibeault C, Obermayer B, Weiner J, Velásquez IM, Gerhard C, Stubbemann P, Hanitsch LG, Pischon T, Witzenrath M, Sander LE, Kurth F, Meyer-Hermann M, de la Rosa K. Soluble ACE2 correlates with severe COVID-19 and can impair antibody responses. iScience. 2024 Feb 24;27(3):109330. doi: 10.1016/j.isci.2024.109330. PMID: 38496296; PMCID: PMC10940809.Soluble ACE2 correlates with severe COVID-19 and can impair antibody responses — PubMed
Li X., Wirtz T., Weber T., Lebedin M., Lowenstein E.D., Sommermann T., Zach A., Yasuda T., de la Rosa K., Chu V.T., Schulte J.H., Müller I., Kocks C., Rajewsky K. Precise CRISPR-Cas9 gene repair in autologous memory T cells to treat familial hemophagocytic lymphohistiocytosis. Sci Immunol 9, eadi0042, 2024. https://doi.org/10.1126/sciimmunol.adi0042
Roske Y, Cappel C, Cremer N, Hoffmann P, Koudelka T, Tholey A, Heinemann U, Daumke O, Damme M. Structural analysis of PLD3 reveals insights into the mechanism of lysosomal 5′ exonuclease-mediated nucleic acid degradation. Nucleic Acids Res. 2024 Jan 11;52(1):370 – 384. doi: 10.1093/nar/gkad1114. PMID: 37994783; PMCID: PMC10783504.Structural analysis of PLD3 reveals insights into the mechanism of lysosomal 5′ exonuclease-mediated nucleic acid degradation — PubMed
Lebedin M., Ratswohl C., Garg A., Schips M., Vazquez Garcia C., Spatt L., Thibeault C., Obermayer B., Weiner J., Moreno Velásquez I., Gerhard C., Stubbemann P., Hanitsch L.G., Pischon T., Witzenrath M., Sander L.E., Kurth F., Meyer-Hermann M., de la Rosa K. Soluble ACE2 correlates with severe COVID-19 and can impair antibody responses. iScience 27, 109330, 2024. https://doi.org/10.1016/j.isci.2024.109330
Zhang J., Sommermann T., Li X., Gieselmann L., de la Rosa K., Stecklum M., Klein F., Kocks C., Rajewsky K. LMP1 and EBNA2 constitute a minimal set of EBV genes for transformation of human B cells. Front Immunol 14: 1331730, 2023. https://doi.org/10.3389/fimmu.2023.1331730
Ratswohl C., Vázquez García C., Ahmad A.U.W., Gonschior H., Lebedin M., Silvis C.E., Spatt L., Gerhard C., Lehmann M., Sander L.E., Kurth F., Olsson S., de la Rosa K. A design strategy to generate a SARS-CoV‑2 RBD vaccine that abrogates ACE2 binding and improves neutralizing antibody responses. Eur J Immunol 53(10): e2350408, 2023. https://doi.org/10.1002/eji.202350408
Secker C., Motzny A.Y., Kostova S., Buntru A., Helmecke L., Reus L., Steinfort R., Brusendorf L., Boddrich A., Neuendorf N., Diez L., Schmieder P., Schulz A., Czekelius C., Wanker E.E. The polyphenol EGCG directly targets intracellular amyloid‑β aggregates and promotes their lysosomal degradation. J Neurochem 166(2): 294 – 317, 2023. https://doi.org/10.1111/jnc.15842
M. Broto, M.M. Kaminski, C. Adrianus, N. Kim, R. Greensmith, S. Dissanayake-Perera, A.J. Schubert, X. Tan, H. Kim, A.S. Dighe, J.J. Collins, and M.M. Stevens (2022) Nanozyme-catalysed CRISPR assay for preamplification-free detection of non-coding RNAs. Nature Nanotechnology, 17, 1120 – 1126.
A.C. McGarvey, W. Kopp, D. Vucicevic, K. Mattonet, R. Kempfer, A. Hirsekorn, I. Bilic, M. Gil, A. Trinks, A.M. Merks, D. Panáková, A. Pombo, A. Akalin, J.P. Junker, D.Y.R. Stainier, D. Garfield, U. Ohler, and S.A. Lacadie (2022) Single-cell-resolved dynamics of chromatin architecture delineate cell and regulatory states in zebrafish embryos. Cell Genomics, 2, 100083.
U. Zinnall, M. Milek, I. Minia, C.H. Vieira-Vieira, S. Müller, G. Mastrobuoni, O.G. Hazapis, S. Del Giudice, D. Schwefel, N. Bley, F. Voigt, J.A. Chao, S. Kempa, S. Hüttelmaier, M. Selbach, and M. Landthaler (2022) HDLBP binds ER-targeted mRNAs by multivalent interactions to promote protein synthesis of transmembrane and secreted proteins. Nature Communications, 13, 2727.
W. Winick-Ng, A. Kukalev, I. Harabula, L. Zea-Redondo, D. Szabó, M. Meijer, L. Serebreni, Y. Zhang, S. Bianco, A.M. Chiariello, I. Irastorza-Azcarate, C.J. Thieme, T.M. Sparks, S. Carvalho, L. Fiorillo, F. Musella, E. Irani, E.T. Triglia, A.A. Kolodziejczyk, A. Abentung, G. Apostolova, E.J. Paul, V. Franke, R. Kempfer, A. Akalin, S.A. Teichmann, G. Dechant, M.A. Ungless, M. Nicodemi, L. Welch, G. Castelo-Branco, and A. Pombo (2021) Cell-type specialization is encoded by specific chromatin topologies. Nature, 599, 684 – 691.
E. Dias D’Andréa, Y. Roske, G.A.P. de Oliveira, N. Cremer, A. Diehl, P. Schmieder, U. Heinemann, H. Oschkinat, and J. Ricardo Pires (2020) Crystal structure of Q4D6Q6, a conserved kinetoplastid-specific protein from Trypanosoma cruzi. Journal of Structural Biology, 211, 107536.
M. Zech, R. Jech, S. Boesch, M. Škorvánek, S. Weber, M. Wagner, C. Zhao, A. Jochim, J. Necpál, Y. Dincer, K. Vill, F. Distelmaier, M. Stoklosa, M. Krenn, S. Grunwald, T. Bock-Bierbaum, A. Fecíková, P. Havránková, J. Roth, I. Príhodová, M. Adamovicová, O. Ulmanová, K. Bechyne, P. Danhofer, B. Veselý, V. Han, P. Pavelekova, Z. Gdovinová, T. Mantel, T. Meindl, A. Sitzberger, S. Schröder, A. Blaschek, T. Roser, M.V. Bonfert, E. Haberlandt, B. Plecko, B. Leineweber, S. Berweck, T. Herberhold, B. Langguth, J. Švantnerová, M. Minár, G.A. Ramos-Rivera, M.H. Wojcik, S. Pajusalu, K. Õunap, U.A. Schatz, L. Pölsler, I. Milenkovic, F. Laccone, V. Pilshofer, R. Colombo, S. Patzer, A. Iuso, J. Vera, M. Troncoso, F. Fang, H. Prokisch, F. Wilbert, M. Eckenweiler, E. Graf, D.S. Westphal, K.M. Riedhammer, T. Brunet, B. Alhaddad, R. Berutti, T.M. Strom, M. Hecht, M. Baumann, M. Wolf, A. Telegrafi, R.E. Person, F.M. Zamora, L.B. Henderson, D. Weise, T. Musacchio, J. Volkmann, A. Szuto, J. Becker, K. Cremer, T. Sycha, F. Zimprich, V. Kraus, C. Makowski, P. Gonzalez-Alegre, T.M. Bardakjian, L.J. Ozelius, A. Vetro, R. Guerrini, E. Maier, I. Borggraefe, A. Kuster, S.B. Wortmann, A. Hackenberg, R. Steinfeld, B. Assmann, C. Staufner, T. Opladen, E. Ružicka, R.D. Cohn, D. Dyment, W.K. Chung, H. Engels, A. Ceballos-Baumann, R. Ploski, O. Daumke, B. Haslinger, V. Mall, K. Oexle, and J. Winkelmann (2020) Monogenic variants in dystonia: an exome-wide sequencing study. Lancet Neurology, 19, 908 – 918.
D. Sundaravinayagam, A. Rahjouei, M. Andreani, D. Tupina, S. Balasubramanian, T. Saha, V. Delgado-Benito, V. Coralluzzo, O. Daumke, and M. Di Virgilio (2019) 53BP1 supports immunoglobulin class switch recombination independently of its DNA double-strand break end protection function. Cell Reports, 28, 1389 – 1399.
H. Behrmann, A. Lürick, A. Kuhlee, H. Kleine Balderhaar, C. Bröcker, D. Kümmel, S. Engelbrecht-Vandré, U. Gohlke, S. Raunser, U. Heinemann, and C. Ungermann (2014) Structural identification of the VPS18 β-propeller reveals a critical role in the HOPS complex stability and function. Journal of Biological Chemistry, 289, 33503 – 33512.
F. Mayr and U. Heinemann (2013) Mechanisms of Lin28-mediated miRNA and mRNA regulation: a structural and functional perspective. International Journal of Molecular Sciences, 14, 16532 – 16553.
S. Boivin and D.J. Hart (2011) Interaction of the influenza A virus polymerase PB2 C‑terminal region with importin a isoforms provides insights into host adaptation and polymerase assembly. Journal of Biological Chemistry, 286, 10439 – 10448.
H. Striegl, M.A. Andrade-Navarro, and U. Heinemann (2010) Armadillo motifs involved in vesicular transport. PLoS ONE, 5, e8991.
News
Transfer & Outreach
On this page, you’ll find information about our transfer and outreach activities such as a structure gallery of proteins and patents.
- Protein Data Bank
Below, you find our crystal structure depositions into the Protein Data Bank.
- Patents
Explore our innovative contributions to the field through our patent portfolio, accessible via the following link: