New brick in PKA Legoland adds complexity to a picture we thought we knew well
Dr Oliver Rocks wasn’t planning to investigate PKA regulation when he stumbled on the protein in a screen for regulators involved in the signalling pathways that control the cell’s cytoskeleton. The cell constantly remodels the cytoskeleton’s network of filaments and the Rocks lab works to understand how these changes are controlled. Finding that PKA was linked to these signalling processes was not entirely unexpected, but the subunit identified in the screen was unusual. “We were surprised to find the catalytic subunit of PKA, because normally control of this pathway is through the regulatory subunit” Rocks says.
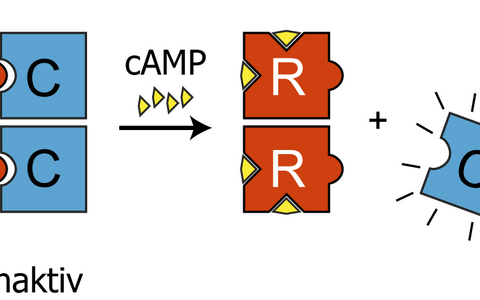
To understand why the catalytic subunit of PKA raised eyebrows we need to join the young biochemists in their lecture theatre. They’re drawing the textbook picture of PKA regulation, which looks like four lego bricks – let’s say two blue and two purple. The blue bricks are the regulatory subunits and their job is to hold onto the purple catalytic subunits and stop them sending signals by phosphorylating proteins. They only release the catalytic subunits when the cell receives a signal that increases the levels of the chemical cAMP. Imagine cAMP as tiny little red lego bricks that clip onto the blue regulatory subunits and force them to set the purple catalytic subunits free.
This lego-brick diagram is what biochemistry students draw in their notepads. The story is of course more complicated, and there are extra layers of regulation in different contexts. But like the classic diagram, the extra layers happen via the regulatory subunits. Surprisingly, it was the catalytic subunits that turned up in Rocks’ screen, so he and his team set out to find out what was happening.
In the screen the catalytic subunit of PKA (PKAC) was binding to a protein called ARHGAP36. It was an uncharacterised protein, which means that no-one knew much about it. ARHGAP36 gets the GAP in its name from its family, the GTPase Activating Proteins (GAPs) which regulate cellular signalling pathways that remodel the cytoskeleton.
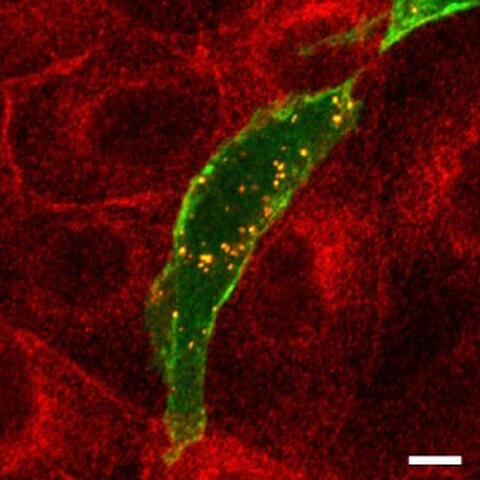
PKAC, fluorescing red, and ARHGAP36, fluorescing green, accumulates in small vesicles (appearing yellow).
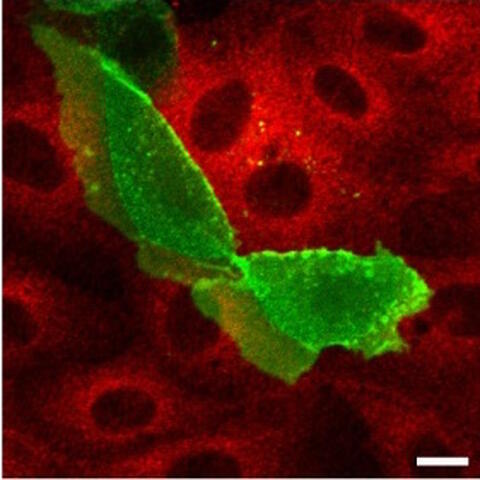
Later, PKAC completely disappears.
Rebecca Eccles investigated how ARHGAP36 interacts with PKAC for her PhD research in the Rocks lab. She worked together with other scientists at the MDC in Berlin, as well as collaborators at the University of Liverpool. Eccles found that ARHGAP36 can turn off PKAC in two ways: By binding to it and blocking its action, and by sending it on the path to one of the cell’s degradation centres.
PKA’s job in the cell is to pass on signals, which the catalytic subunit does by its kinase action – kinases attach a phosphate molecule to their target proteins (substrates). ARHGAP36 stops PKAC from binding its substrates a bit like a key stuck in a lock stops you from opening a door. If we imagine PKAC as a lock and its substrates as keys, then we can imagine ARHGAP36 as a skeleton key –a pseudosubstrate. It fits into PKAC’s active site and sits there, blocking substrates from binding.
Looking at cells expressing green-labelled ARHGAP36, Eccles found that ARHGAP36 wasn’t just deactivating PKAC, it was also causing red-labelled PKAC to disappear. After a few hours the two proteins moved together into vesicles in the cell, and later the PKAC completely disappeared.
PKAC was being tagged with ubiquitin, a label that tells cells a protein needs to be broken down. Then it was being degraded in the lysosome. “It was a very strong downregulation and since PKAC is a cytosolic protein we were surprised at it being degraded by the endolysosomal pathway,” Eccles says. Most cytosolic proteins are chewed up by a cellular degradation machine called the proteasome, whereas the endolysosome pathway normally breaks down membrane proteins.
Since PKA is found in diverse tissues, the scientists wanted to find out where and when ARHGAP36 inhibits it. “ARGAP36 is a strong inhibitor, so you wouldn’t want it turning off PKA everywhere,” Rocks explains. They found that ARHGAP36 is not present in all cells all the time – in fact its expression is quite limited. It’s turned on during embryonic muscle development, perhaps because keeping PKA in check is important in this context. ARHGAP36 is also upregulated in at least one of the four subtypes of medulloblastoma, the most common childhood brain cancer, as well as in neuroblastoma, another frequent cancer in young children. “From a cancer point of view, it could be that tight regulation is critical so that PKA is not deactivated where it’s needed,” Rocks says. Other scientists have recently reported that changes to PKA signalling may influence tumour progression in many cancer types.
Finding that PKA, one of the best studied proteins, can be regulated in an unknown way is intriguing. The exact biological role of ARHGAP36 remains to be discovered, but it may well play a role in muscle development and tumour progression for some cancers. Understanding how signalling pathways are regulated may also be useful in future drug development: indirectly regulating a proteins levels provides further options for enzymes that are otherwise hard to manipulate. In any case, the studies of ARHGAP36 in the Rock’s lab have added new complexity to the classic lego-brick understanding of PKA regulation.
Rebecca L. Eccles1, Maciej T. Czajkowski1,2,Carolin Barth1, Paul Markus Müller1, Erik McShane1, Stephan Grunwald1, Patrick Beaudette1, Nora Mecklenburg1, Rudolf Volkmer4, Kerstin Zühlke1, Gunnar Dittmar1, Matthias Selbach1, Annette Hammes1, Oliver Daumke1,5, Enno Klussmann1,6, Sylvie Urbé3, Oliver Rocks1 (2016): “Bimodal antagonism of PKA signalling by ARHGAP36.” Nature Communications. doi:10.1038/NCOMMS12963
1Max Delbrück Center for Molecular Medicine in the Helmholtz Association, Berlin, Germany; 2Berlin Institute of Health (BIH), Berlin, Germany; 3Cellular and Molecular Physiology, Institute of Translational Medicine, University of Liverpool, Liverpool, UK; 4Leibniz-Institut für Molekulare Pharmakologie, Berlin, Germany; 5Institute for Chemistry and Biochemistry, Freie Universität Berlin, Berlin, Germany; 6DZHK, German Centre for Cardiovascular Research, Berlin, Germany
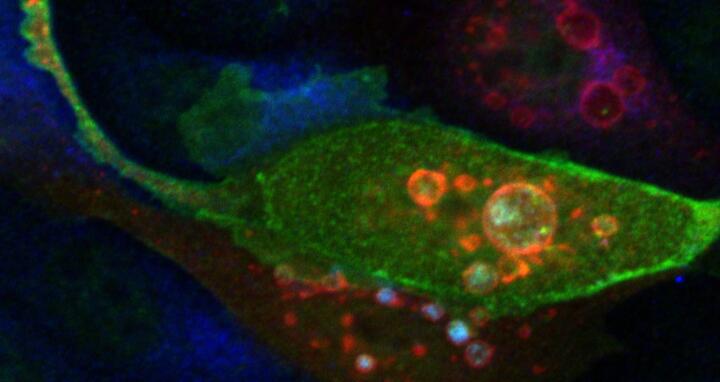
Featured Image: PKAC, fluorescing blue, accumulates inside red enlarged vesicles when ARHGAP36, fluorescing green, is present. Image: Rebecca Eccles, MDC.