Perfecting the tools to rewrite genomes
For many years, scientists have dreamed of manipulating DNA the way copy-editors change texts: from replacing single letters of the genetic code to altering sequences of any length and adding, deleting, replacing, or rearranging large regions of chromosomes. Changes of all of these types have occurred naturally over the course of evolution, and they even occur when DNA from two parents is combined to create a new genome. Recent years have seen the development of very powerful new methods to delete sequences from a chromosome and insert new ones, but they aren’t efficient enough to easily rewrite single letters of the code.
Now the groups of Klaus Rajewsky and Ralf Kühn at the MDC have improved a method of genetic engineering known as CRISPR/Cas. This system has already been used for research to alter all types of cells – including fertilized eggs – but the new modifications represent a method for introducing precise mutations into genomes at a high frequency and selectivity. The work, which was published online by Nature Biotechnology on March 24, promises to have a profound effect on the study of diseases and, potentially, on the development of new forms of therapies.
Background
Like many of the tools of genetic engineering, CRISPR/Cas is based on molecules originally found in bacteria, which scientists have adapted for use in mice and other experimental organisms. Originally bacteria evolved this system as a means of protecting themselves from viruses, which often introduce short sequences of their own DNA into the bacterial genomes during an infection.
Successful insertions usually create sequences called spacer regions between protein-coding genes. These regions often consist of many small, repeated sequences that are the source of the name CRISPR (for clustered regularly interspaced short palindromic repeats). Bacterial molecules called CRISPR-associated genes, or Cas, recognize CRISPR sequences and cut both strands of DNA. Cas proteins are guided to selected target sites by short guiding (sg) RNAs with specific sequences.
Double-stranded breaks in DNA occur naturally, and are also produced by exposure to radiation or DNA-damaging chemicals. When such breaks occur in an organism’s genome, they must be repaired or the cell will die. This phenomenon is the basis of radio- and chemotherapies for cancer. Cells that reproduce quickly (such as those of a tumor) often don’t have enough time to make the repairs before they die.
Cells engage in two basic types of repair: homologous and non-homologous. The first type is homology-directed (HD) repair, which is much more rare in mammalian cells. It depends on the presence of an exact match between the sequences at the ends of a “broken” fragment and an existing, homologous (or “matching”) sequence in a genome. If such a match is found, enzymes that mediate the recombination of DNA molecules may insert a new gene segment, placed between two homology regions of an artificial DNA molecule that is used as a template for the repairs.
HD repair allows scientists to plan and carry out precise mutations in genomes, but this occurs only at low frequency in mammalian cells. The other type of repair, called the non-homologous end-joining (NHEJ) repair pathway, is much more efficient in mammalian cells. But it is also less precise, since it simply reconnects free DNA ends without a repair template. Often nucleotides are lost or gained at random in the process. This means that NHEJ repair can be used to create short deletions in sequences, but they are not precise enough to permit modifications such as the insertion or rewriting of specific gene segments. The prevalence of NHEJ repair has been a central limitation in the types of changes genetic engineers have been able to make in genomes.
The current project
Mice and other multicellular organisms generally don’t have Cas proteins that cut the DNA double strand. But within the past few years scientists have learned to introduce them artificially as a means of interfering with specific genes. This is generally done by attaching Cas to a single guide RNA (sgRNA), whose sequence matches a specific region of DNA. This tells the cell where to deliver Cas in the genome and make a cut.
So combinations of sgRNAs and Cas can cut a sequence and powerfully interfere with a gene, but the non-homologous repair system has limited both the types and accuracy of alterations that can be made. Klaus Rajewsky, Ralf Kühn, their colleagues at the MDC, and other groups recently hit upon an idea that could change this situation. In fact, the current paper was published back-to-back with a study from Klaus’ former colleague Hidde Ploegh, of the Whitehead Institute and MIT in Massachussetts (USA), which used a comparable approach to accomplish much the same thing. As each group learned of the other’s work, the teams of Klaus, Ralf, and Ploegh agreed to coordinate the publication of their results, which mutually confirmed each other.
“Cells must repair double-stranded breaks in their DNA to survive,” Klaus says. “If they can’t carry out non-homologous repair, then they have to switch to the homologous type or die. So if we could somehow block the non-homologous repair pathway, we might ensure that sgRNAs would deliver Cas to a precise position, and make exactly the type of change we desire.”
The method to accomplish this depends on the fact that homologous and non-homologous repair rely on different enzymes. A protein called DNA ligase IV is essential for – and exclusively involved in – non-homologous repair. By blocking it, the scientists ought to be able to force cells to take the other route.
Several methods were available: the team could “knock down” DNA ligase IV, block its enzymatic activity using a small-molecule inhibitor called SCR7, or express proteins that mark Ligase IV for degradation. The first and second methods led to a four-to-five-fold increase in the amount of homologous repair in cells. The third turned out to be even more effective. By causing cells to express two proteins called E1B55K and E4orf6, which lead to the destruction of DNA ligase IV, the scientists achieved an up to eight-fold increase in homologous repair. Interestingly, these two molecules are derived from a pathogen called the adenovirus, which evolved them as a means of overcoming cellular defenses. The adenovirus introduces E1B55K and E4orf6 molecules into cells; they degrade DNA ligase IV. By adding these molecules to the vector that was used to deliver Cas into cells, the scientists developed a multi-purpose tool that ought to ensure homologous repair.
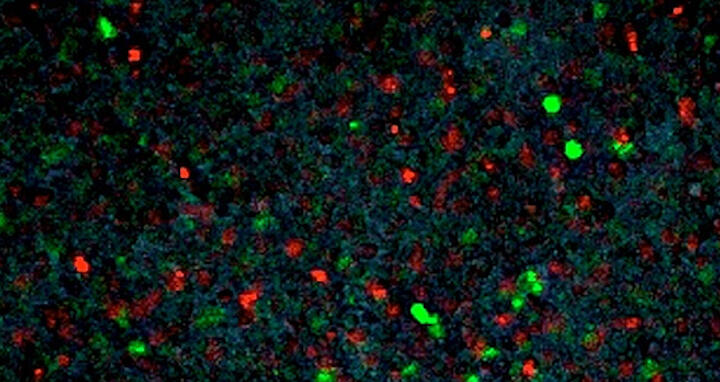
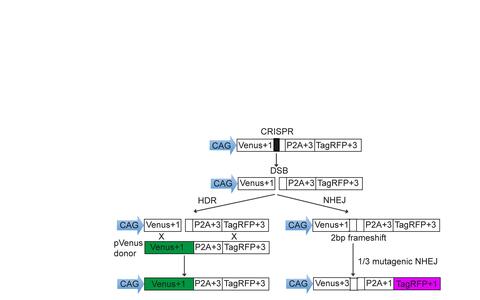
Van Trung Chu, a postdoctoral fellow in Klaus’ group, began a series of experiments to test whether vectors outfitted with sgRNA, Cas, and the adenovirus molecules could deliver a specific sequence to a precise position in the genome of human cells raised in cell cultures. Van Trung introduced a DNA repair “traffic light” reporter system that would give off green fluorescent signal if the construct underwent HD repair, and a red signal if the NHEJ pathway was used.
The experiment passed both tests. The team had demonstrated that they could precisely and selectively target a region of DNA and introduce whatever type of change they liked through HDR. NHEJ repair, as indicated by the loss of red signals, was almost completely abolished.
Could the method also be used to develop new genetically modified organisms such as mice? Traditionally, the development of such animals by introducing changes in embryonic stem cells has required at least a year, sometimes two. Klaus turned to Ralf Kühn, who had been a member of his lab in Cologne a decade ago. After a stint in Munich, Ralf recently established his own group at MDC. He brings along a great deal of experience in reprogramming stem cells and introducing genetic modifications that can be used to study diseases. He is also head of the Transgenic Core Facility, which helps scientists develop new animal models. The facility is also an important component of the Berlin Institute of Health, where it is serving collaborative projects of the MDC and Charité.
The MDC facility, like many others in the world, had already successfully used Cas and sgRNAs to directly introduce mutations into fertilized mouse eggs. The new system does not require culturing embryonic stem cells and saves an enormous amount of time by reducing these steps to a single intervention in one-cell embryos. But as in mouse or human cell lines, NHEJ is the dominating repair mechanism in one-celled embryos, which has restricted the number of precisely modified mutants obtained in any one experiment. The problem could be solved by exposing manipulated embryos to the small molecule inhibitor of Ligase IV, as shown by the team of H. Ploegh. Presently the MDC facility is testing whether the use of adenovirus proteins may yield even more precise mutants, using a traffic light reporter mouse line they developed.
Broader implications
Scientists at the MDC and elsewhere are already employing the CRISPR/Cas system to develop new lines of cells and organisms for disease research. They can now use it to imitate mutations and other genetic changes that are involved in disease much more easily, study their biological effects, and try out new forms of treatment.
At the same time, Klaus points out, the work has much broader implications that immediately attracted the attention of researchers and the media. A central issue concerns the potential of genetic engineering to make changes in human germ line cells. This idea evokes a number of ethical and moral concerns that have been voiced for many years. Until now, technical limitations in “genome editing” have posed significant barriers to using the methods to alter human germ line cells and, for example, correct hereditary defects in genes. The rate of success has been comparatively low due to the prevalence of non-homologous repair in mammalian cells. In other words, alterations might be fatal to the cells, or produce additional, undesirable defects. With the improvements, which will surely be rendered even more effective over time, the correction of defects will likely become highly reliable and efficient. While introducing such changes might be desirable for many families affected by devastating diseases, the same methods could be used to introduce other types of changes in a human embryo.
In mid-March, the publication of several related papers on CRISPR/Cas technology demonstrated that many of the technical obstacles are likely to be resolved very soon. On March 19, David Baltimore, Keith Yamamoto, and 16 other prominent scientists published an urgent appeal on the matter in the journal Science. The letter discouraged any modifications of human germ line cells – even for clinical applications deemed urgent – while the “societal, environmental, and ethical implications of such activity are discussed among scientific and governmental organizations.” It stressed the need for increased educational programs and forums for public discussion, for transparent research to study the precision of the method, and the creation of a global group of experts in all relevant areas to consider the implications of manipulations of the human genome. Klaus and his colleagues will certainly play an active role in these discussions.
In the meantime, Klaus says, the improved system offers life scientists their most powerful tool to date to investigate the biology of hereditary diseases in cells and model organisms. It will reduce the number of animals needed for biomedical research and surely deliver more precise insights into many diseases that afflict humanity.
Reference:
Van Trung Chu, Timm Weber, Benedikt Wefers, Wolfgang Wurst, Sandrine Sander, Klaus Rajewsky, Ralf Kühn. Increasing the efficiency of homology-directed repair for CRISPR-Cas9-induced precise gene editing in mammalian cells. Nature Biotechnology (2015), doi:10.1038/nbt.3198
Further information:
Ethics of embryo editing divides scientists, Nature (18 March 2015)
Featured Image: This image shows cells that have been transfected with the “stoplight” reporter module. If the sequence has been placed into the genome using the non-homologous end joining (NHEJ) pathway, then sequences are disrupted and the cell gives off a red signal. If homologous repair has been used, the sequence remains intact and cells insert the probe in between the breaks, giving off a green signal. The image is typical for previous experiments using CRISPR/Cas, when both pathways were active in cells. The new experiments show that blocking the NHEJ pathway leads to homologous repair and most cells emit the green signal Photo: Lab Ralf Kühn, MDC