Unwinding defective messenger RNAs
Evolution has produced quality control systems that permit our cells to detect and destroy defective messenger RNAs before they go on to produce faulty proteins. Markus Landthaler’s group has now discovered how a protein called an RNA helicase helps process RNAs so that they can be destroyed or recycled. The work appeared in the May 22 edition of Molecular Cell.
The chemistry of life is imperfect. Over the long term, this has led to the evolution of humans and every other species; over the short term, it usually causes disease. For individuals, mutations and other types of changes in DNA are typically harmful if they affect genes and alter the recipes of the messenger RNAs encoded in them. Faulty RNA molecules can also arise through simple errors as they are manufactured by cells. Such molecules may go on to create defective proteins that can’t carry out essential functions in our cells. But evolution has produced quality control systems that often catch and destroy defective RNAs before they lead to harmful proteins. Markus Landthaler’s group has now discovered how a protein helps process RNAs so that they can be destroyed. The work appeared in the May 22 edition of Molecular Cell.
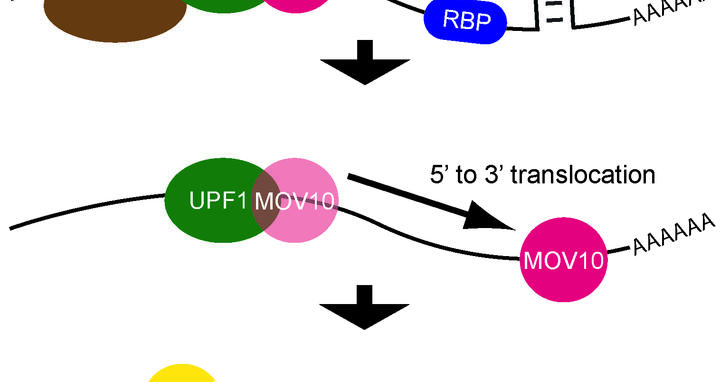
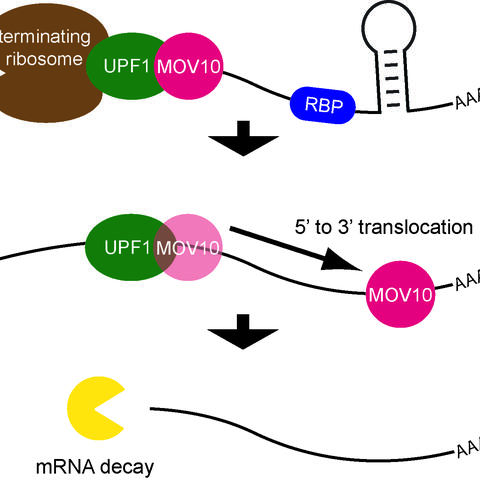
A messenger RNA (mRNA) is a string-like molecule with several regions that play an important role in its translation into proteins. It begins with a five-prime (5’) region, followed by a central coding region, and then a three-prime (3’) tail. Proteins dock onto the 5’ region and build a platform for a molecular machine called the ribosome. This machine travels down the molecule, reading the information in the coding region and using it as a set of instructions to assemble a protein. The ribosome stops at a sequence called a termination codon. The 3’ tail provides docking sites for proteins that influence this process of translation, sometimes blocking it entirely or causing the RNA to be recycled.
One type of protein that may dock on is a RNA helicase, which can change an RNA’s structure and remove proteins that bind to it. RNA helicases unwind the RNA and change the regions that are exposed, which facilitates mRNA decay. One such helicase is UPF1, which participates in a process called nonsense-mediated mRNA decay, or NMD, which breaks down defective RNAs. NMD recognizes that the mRNA has a termination codon too early because of a mutation or another defect, which would usually lead to a shortened protein that might not be able carry out important cellular functions. In the current study, Lea Gregersen and other members of the lab examined the behavior of an RNA helicase called MOV10. “The way this protein interacts with RNAs hadn’t been studied,” Markus says. “We knew that it bears similarities to helicases. We also knew that it blocks the activity of two viruses, HIV1 and hepatitis C, which replicate by fooling infected cells into producing viral RNAs. MOV10 probably helps the cell recognize that there is something wrong with these molecules.”
Understanding the function of an RNA helicase such as MOV10 requires identifying which RNAs it binds to and the precise sites in molecules where it attaches itself, and then determining exactly what it does to its RNA targets. To understand the normal activity of MOV10, the scientists compared it with two mutant forms that are unable to function as helicases. To find out where the helicase attached itself, Lea and her colleagues used a method called PAR-CLIP. This replaces some of the subunits of RNA with altered versions; when exposed to ultraviolet light, the helicase gets locked onto the RNA in a specific position. The scientists purified the MOV10 and the target transcripts it was bound to, sequenced the RNAs with the help of deep sequencing from Wei Chen’s lab, and found that MOV10 can bind to about 12,500 different protein-encoding mRNAs.
Next the scientists discovered that the helicase binds to mRNAs in a region of the 3’ tail that directly follows the termination codon. It then travels downstream along the mRNA farther toward the 3’ end, unwinding the transcript in the process. This changes the molecule’s structure and displaces proteins that would otherwise bind to RNA and make it accessible to molecules involved in RNA degradation.
“When we studied the mutant forms of the helicase, we found that they could bind to targets but were unable to unwind them,” Markus says. “So they were unable to make changes in RNA that would otherwise trigger its degradation.”
The lab discovered that MOV10 binds directly to UPF1, which plays a key role in nonsense-mediated mRNA decay and in the degradation of mRNAs with long 3’ tails. Another experiment that eliminated MOV10 from cells led to target RNAs that survived longer and produced more proteins – probably because mRNA decay is slowed down.
“The study gives us deeper insights into a process that is essential in the quality control of cellular molecules,” Markus says. “We think that by unwinding RNA, MOV10 and other helicases alter molecular structures, remove other proteins, and allow RNAs to be targeted for degradation. Our work shows that PAR-CLIP can be applied to other RNA-binding proteins with deficiencies in their enzymatic activities to study how these interactions regulate their translation into proteins, their turnover, and other crucial processes.”
Featured Image: On the trail of the chemistry of life: in the current project, Markus Landthaler and his lab investigated cellular mechanisms involved in the detection of faulty messenger RNAs. Photo: David Ausserhofer, MDC
Highlight Reference:
Gregersen LH, Schueler M, Munschauer M, Mastrobuoni G, Chen W, Kempa S, Dieterich C, Landthaler M. MOV10 Is a 5′ to 3′ RNA Helicase Contributing to UPF1 mRNA Target Degradation by Translocation along 3′ UTRs. Mol Cell. 2014 May 22;54(4):573 – 85. doi: 10.1016/j.molcel.2014.03.017.
Full text of the original article