Glues, chains, and misfolded proteins
Proteins need to have a precise structure to carry out their functions in cells. As they are produced, they undergo a complex process of folding that gives them the proper shape, arranging their chemical subunits (amino acids) in a way that allows them to bind to specific molecules and carry out their tasks. But no biological system is perfect, and sometimes cells produce improperly folded molecules. They may be unable to dock onto their partners, or bind to the wrong ones. This can lead to serious diseases, mainly because improperly folded proteins tend to clump together and disrupt cellular processes – found, for example, in many neurodegenerative diseases. Cells have intricate mechanisms to recognize such proteins and destroy them before they can do any harm. One of the major systems involves attaching a small protein called ubiquitin to misfolded proteins, either singly or in long chains. This activates a mechanism that destroys the molecules – a discovery that led to the 2004 Nobel Prize in Chemistry for Israeli researcher Aaron Ciechanover. Now Thomas Sommer’s lab, in collaboration with Ciechanover, has discovered how a protein called Cue1p helps attach long ubiquitin chains to misfolded molecules. The study appeared in the May 23 issue of Molecular Cell.
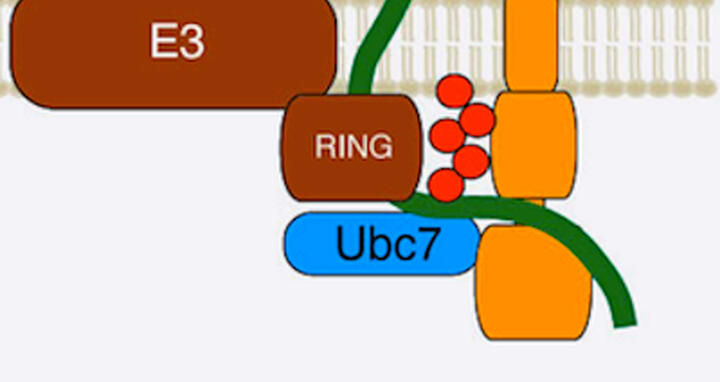
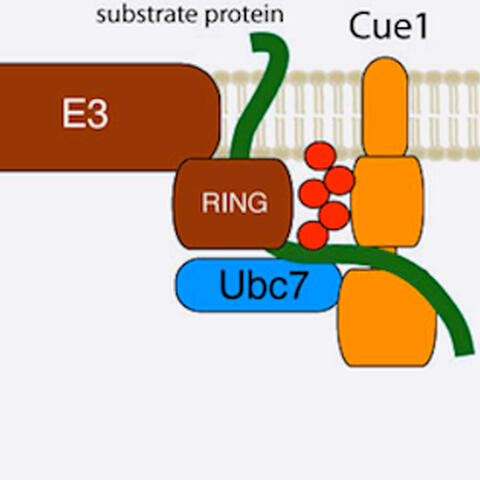
The current study shows that Cue1 attaches long chains of ubiquitin molecules (red) to a misfolded protein (green) as it is translated. The cell then recognizes that the molecule should be destroyed.
Ubiquitin is found in all eukaryotic cells – a branch of life including all animals as well as single-celled organisms such as yeast. The molecule arose long ago in simple organisms and has such important functions that it has been conserved throughout the evolution of more complex forms of life. Similarly, the enzymes that attach ubiquitins to misfolded proteins have been conserved. This has allowed researchers to use model organisms such as yeast to study the mechanisms of ubiquitylation and the way cells destroy proteins.
The chemical properties of ubiquitin allow it to be attached to an amino acid called lysine in target proteins. Sometimes the attachment of a single ubiquitin molecule leads to the destruction of the target. But ubiquitin itself also has seven lysine building blocks, which means that more copies can be attached onto the small molecule. This can lead to the formation of long chains of ubiquitin.
“You can think of the attachment of ubiquitin like a label that is added to its target molecule,” Thomas says. “To read this label, nature has evolved so-called ubiquitin-binding domains (UBDs), which recognize specific kinds of modifications that ubiquitin has made to proteins.”
In the current study, Katrin Bagola and other members of Thomas’ lab investigated a protein called Cue1p in yeast. The scientists knew that this molecule was required to attract a molecule called Ubc7p, which glues ubiquitin onto defective proteins that are mounted in cell membranes. Without Cue1p, Ubc7p was not functional.
Cue1p has a ubiquitin-binding domain called the CUE domain. Thomas and his co-workers made the surprising discovery that this domain of Cue1p only binds to chains containing multiple ubiqitins. “We had always been frustrated in our attempts to show that Cue1p could bind to ubiquitin,” Thomas says. “But we’d been expecting it to bind to single copies of the molecule; now we discover that it only binds to polyubiquitin chains.”
The scientists began creating mutant forms of Cue1p and introducing them into yeast cells. The goal was to determine if the CUE domain of the protein directly binds to ubiquitin – or whether it requires some other molecule as a “mediator” – and, if so, what parts of its architecture are responsible.
Katrin and her colleagues discovered that Cue1p binds to growing ubiquitin chains and thereby strongly promotes the lengthening of these chains. Such long chains are necessary for the destruction of many misfolded membrane proteins.
“This gives us insights into an important mechanism for the destruction of defective proteins in yeast that may also be relevant for other organisms,” Thomas says. “That’s significant because defects in this process have been linked to a number of serious diseases. But it also shows that ubiquitin-binding regions such as CUE may have multiple functions. In this case, for example, the region recognizes and helps extend chains of multiple ubiquitin molecules – so it helps us understand more general principles of the formation of long ubiquitin chains.”
- Russ Hodge
Highlight Reference:
Bagola K, von Delbrück M, Dittmar G, Scheffner M, Ziv I, Glickman MH, Ciechanover A, Sommer T. Ubiquitin Binding by a CUE Domain Regulates Ubiquitin Chain Formation by ERAD E3 Ligases. Mol Cell. 2013 May 23;50(4):528 – 39.
Wikipedia article on ubiquitin
Aaron Ciechanover’s Nobel prize lecture on protein degradation