ERC Starting Grant awarded to Fabian Coscia
The junior research group led by Dr. Fabian Coscia, set up in 2021, is getting a boost from an ERC Starting Grant. In addition to prestige, the grant provides €1.5 million in funding over five years while also opening doors to other opportunities and attracting high-quality applicants for doctoral and postdoc positions.
“This is a huge recognition for our work, and I’m very excited about the funding for the next several years,” says Coscia. “We’re currently establishing methods for performing high-resolution spatial proteomics. After that we plan to apply them to complex tumor tissue – where we are among the first to do so.” The researchers hope, among other things, to use the cutting-edge technologies to uncover novel mechanisms by which cancer cells can evade therapies and thus ensure their survival.
The international peer reviewers of the ERC look for novel approaches that could open up new frontiers and spur significant advances (“high risk, high reward” research). The researchers must have completed their PhD within the last two to seven years and have a scientific track record showing great promise. This year, 400 European scientists from a wide range of disciplines will receive ERC Starting Grants.
The Deep Spatial Proteomics project, in detail
When their cancer returns after a first round of treatment, patients with squamous cell carcinoma of the head and neck often put much hope in currently available immunotherapies. But such treatment helps only about two out of ten of these patients. Coscia and his colleagues want to learn which cells drive the disease, how to recognize them in clinical practice and how to ultimately keep them in check by using targeted drugs.
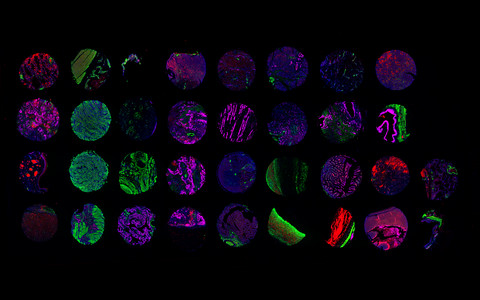
Each image section shows a tissue sample collected from patients. The different cell types were visualized using multiplex immunofluorescence staining.
The local cellular neighborhood – or microenvironment of tumor cells – plays a critical role. While some cells thwart the aggressive behavior of neighboring cells, others help them to spread through the body. Coscia believes this dynamic interplay is inadequately captured if the sole focus is on genes. “What influence genetic changes have on the signaling pathways in the cell and in a tissue depends on whether, and to what extent, the cell actually produces a mutated protein.”
Observing this event at very high resolution for the entirety of the proteins produced (proteome) was previously impossible. “Great progress is now being made in spatial single-cell proteomics based on ultra-sensitive mass spectrometry, an approach that we have been co-developing,” says Coscia. “Using these methods to analyze tissue samples is the next big step in our efforts to decipher the tumor proteome.”
A journey into the past – to create tomorrow’s medicine
We can use our multilayered data to determine which cells and, more importantly, which proteins play a key role when therapy resistance occurs.
Coscia’s group is collaborating closely with Charité – Universitätsmedizin Berlin within the MSTARS consortium. As part of the Deep Spatial Proteomics project, the researchers want to retrospectively analyze samples from a cohort of patients with squamous cell carcinoma of the head and neck. The first step involves using high-resolution microscopy to visualize the cell neighborhoods where particularly aggressive cancer cells reside. AI-guided image analysis tracks down any suspicious cells, which are then isolated with utmost precision using laser microdissection. For each of these cells, the researchers analyze several thousand proteins – and leverage machine learning to evaluate the vast amounts of data and connect them to the clinical picture. “It’s kind of like traveling back in time,” explains Coscia. “We know the entire course of the disease, so we know how patients have fared. This allows us to use our multilayered data to determine which cells and, more importantly, which proteins play a key role when therapy resistance occurs.”
In the future, his research group also wants to apply this technology prospectively – and, if possible, find a signature consisting of 5 to 10 proteins in particularly aggressive cells that clinicians can use to select the most promising therapy. “If this works, the technology is not only interesting for cancer,” says Coscia. “It can be applied to all kinds of diseases.”
Further information
- The key to the right treatment
- Protein maps of tumors
- Fabian Coscia seeks to shed new light on cancer cells
- ERC-Grants at the Max Delbrück Center
Contact
Jana Schlütter
Editor, Communications
Max Delbrück Center +49 (0) 30 9406 2121
jana.schluetter@mdc-berlin.de oder presse@mdc-berlin.de
- Max Delbrück Center
-
The Max Delbrück Center for Molecular Medicine in the Helmholtz Association (Max Delbrück Center) is one of the world’s leading biomedical research institutions. Max Delbrück, a Berlin native, was a Nobel laureate and one of the founders of molecular biology. At the locations in Berlin-Buch and Mitte, researchers from some 70 countries study human biology – investigating the foundations of life from its most elementary building blocks to systems-wide mechanisms. By understanding what regulates or disrupts the dynamic equilibrium of a cell, an organ, or the entire body, we can prevent diseases, diagnose them earlier, and stop their progression with tailored therapies. Patients should be able to benefit as soon as possible from basic research discoveries. This is why the Max Delbrück Center supports spin-off creation and participates in collaborative networks. It works in close partnership with Charité – Universitätsmedizin Berlin in the jointly-run Experimental and Clinical Research Center (ECRC), the Berlin Institute of Health (BIH) at Charité, and the German Center for Cardiovascular Research (DZHK). Founded in 1992, the Max Delbrück Center today employs 1,800 people and is 90 percent funded by the German federal government and 10 percent by the State of Berlin.