Predicting the onset and course of Huntington’s disease
“What are the particles that are toxic to the brain?” This is an unanswered question that lies at the heart of many neurodegenerative diseases, explains Professor Erich Wanker. In Huntington’s disease, for example, protein clusters in the nerve cells of patients form deposits. But why these are harmful, what exactly triggers the disease and what the molecular trigger consists of remain basically unknown.
Our results are the best indication to date that seeding-competent huntingtin fibers are actually responsible for the disorder
Together with his team, Prof. Wanker of the Max Delbrück Center for Molecular Medicine (MDC) in Berlin has identified very small precursors of these deposits: protein chains that clump together to form small fibers. Unlike the already known and relatively large protein deposits, these fibers are about ten thousandths of a millimeter long and consist of mutated huntingtin proteins that are attached to one another. Their properties allow researchers to predict the onset and course of the disease. The results have been published in the scientific journal Molecular Cell.
Enables prediction months prior to disease onset
Anne Ast, a doctoral student in Erich Wanker’s team, found the minute huntingtin fibers in the brains of deceased patients, but also in fruit flies, worms, and mice whose cells produced the mutated protein. These fibers formed in mice before larger protein deposits formed and long before the onset of the disease, enabling the scientist to predict months in advance whether an animal would fall ill.
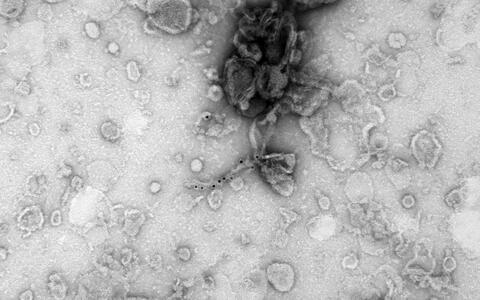
Huntingtin filaments under the electron microscope
The researchers observed the formation of the fibers in a test tube experiment. They fused the mutated huntingtin protein with a fluorescent protein and introduced this artificially produced protein into extracted cerebral tissue. The fluorescent proteins attached themselves to the huntingtin protein chains in the brain extract, elongating the fibrous strand. They were easily detected through their fluorescence.
The research team used this method to determine how fast the harmful fibers would grow and how great their potential was to trigger the formation of new protein chains. After specifically triggering the formation of fibers in the cells, they were able to measure how quickly the fibers formed and thus predict the severity of Huntington’s disease in the genetically modified worms, flies, and mice: The stronger the activity of the particles, the stronger the animals were affected.
Best indication to date
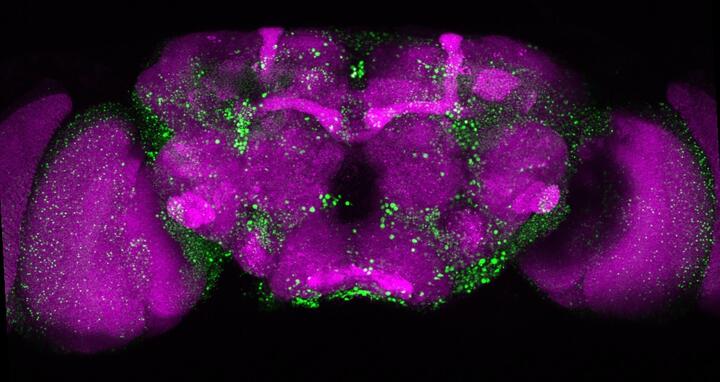
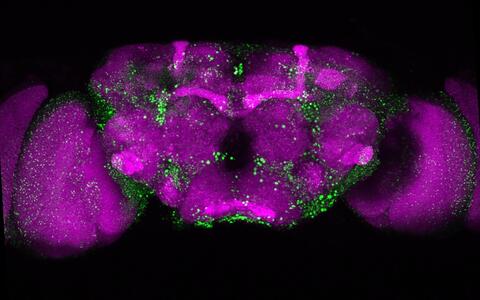
Microscopic image of a fly’s brain. The huntingtin deposits that form as the result of the mutated huntingtin protein are shown in green
“Our results are the best indication to date that seeding-competent huntingtin fibers are actually responsible for the disorder,” says Wanker. His team’s methods, he explains, have rendered something visible that could not be seen before. “We believe this is the best evidence so far that the misfolded protein also triggers Huntington’s,” Wanker continues, explaining that it makes the hypothesis that protein deposits are merely a waste product of the disease even less probable. “This is a huge step for the field.”
Improved research
One of the areas of application for these results would be diagnosis. Wanker explains how, currently, a gene analysis can confirm that a patient will definitely develop Huntington’s disease and die prematurely. However, he continues, being able to provide additional information as to whether the disease will emerge at the age of 40 or 60, or details as to how severe it will be, can help patients cope with the disease. “It would be amazing if we could one day determine the disease progression,” says Wanker. However, the test is still a very long way from having any clinical application.
At the moment, the new method has great potential for research and the search for new drugs. The mutated huntingtin proteins can be extracted from tissue in their earlier stage and tested with pharmaceutical substances to try and inhibit their harmful activity. Wanker hopes that this will lead to the discovery of new agents that could be used to combat the cause of Huntington’s disease.
Literature
Anne Ast, et al. (2018): “mHTT Seeding Activity: A Marker of Disease Progression and Neurotoxicity in Models of Huntington’s Disease.” Molecular Cell. doi: 10.1016/j.molcel.2018.07.032
- The Max Delbrück Center for Molecular Medicine (MDC)
-
The Max Delbrück Center for Molecular Medicine in the Helmholtz Association (MDC) is one of the world’s leading biomedical research institutions. Max Delbrück, a Berlin native, was a Nobel laureate and one of the founders of molecular biology. At the MDC’s locations in Berlin-Buch and Mitte, researchers from some 60 countries analyze the human system – investigating the biological foundations of life from its most elementary building blocks to systems-wide mechanisms. By understanding what regulates or disrupts the dynamic equilibrium in a cell, an organ, or the entire body, we can prevent diseases, diagnose them earlier, and stop their progression with tailored therapies. Patients should benefit as soon as possible from basic research discoveries. The MDC therefore supports spin-off creation and participates in collaborative networks. It works in close partnership with Charité – Universitätsmedizin Berlin in the jointly run Experimental and Clinical Research Center (ECRC), the Berlin Institute of Health (BIH) at Charité, and the German Center for Cardiovascular Research (DZHK). Founded in 1992, the MDC today employs 1,600 people and is funded 90 percent by the German federal government and 10 percent by the State of Berlin.